Flip-Flopping Gene Expression Can Be Advantageous

One gene for pea pod color generates green pods while a variant of that gene gives rise to the yellow-pod phenotype, a feature that helped Gregor Mendel, the 19th century Austrian priest and scientist, first describe genetic inheritance. However, many modern-day geneticists are focused on the strange ability of some genes to be expressed spontaneously in either of two possible ways.
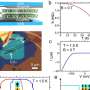
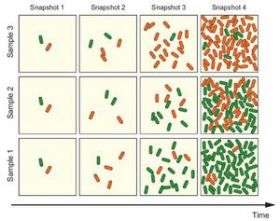
In order to better understand this phenomenon of epigenetic multistability, a major complication for Mendelian genetics, scientists at UC San Diego grew virtual bacterial cells in a computer experiment. They created a two-phenotype model system programmed to grow in ways that matched natural growth. In a deceptively simple experiment, they then recorded the degree to which the two phenotypes varied over time in individual cells, and then repeated the experiment over and over. They reported in the Nov. 19 online edition of Proceedings of the National Academy of Sciences (PNAS) that variability due to epigenetic multistability is larger and persists much longer than they had expected.
While the phenomenon is yet to be discovered in the human genome, the new results suggest that researchers studying bacteria should carefully design their experiments to measure variability due to epigenetic multistability. Even in human cells, multistability may play a role in genes can alternate between “on” and “off” settings.
“Scientists studying bacteria have simply not had the tools to understand phenotypic variability,” said Ting Lu, lead author of the study who was a physics graduate student in the lab of bioengineering professor Jeff Hasty. (Lu is currently a postdoctoral fellow at Princeton University.) “We’ve arrived at a theoretical framework that allows experimenters to measure the ephemeral nature of epigenetics.”
Epigenetic multistability may be vital to cells that are outwardly different, but genetically identical. Only one of the two phenotypes might thrive in a given environmental condition; however, the less advantageous phenotype could come in handy if the environmental conditions unexpectedly changed. By having both phenotypes, the chances would be better that the best one will be present when needed.
Researchers in many labs recently have demonstrated that gene expression can be surprisingly random. The framework established in the UCSD study evaluates how such noisy gene expression affects the properties of developing cellular colonies, such as the progression of bacterial infections or the growth of a population of cancerous cells.
The PNAS paper also documented that the time it takes for a population of cells to reach a stable variance depends on the initial growth conditions and how easily those conditions support cell growth.
“Different results can emerge depending on how cells are grown,” said Jeff Hasty, a professor of bioengineering at UCSD and senior author of the paper. “Our results should allow all researchers studying bacteria to better understand the variability they routinely see from one experiment to the next."
This work is supported by the National Institutes of Health.
Source: UCSD