Researchers make long DNA 'nanowires' for future medical and electronic devices

Ohio State University researchers have invented a process for uncoiling long strands of DNA and forming them into precise patterns. Ultimately, these DNA strands could act as wires in biologically based electronics and medical devices, said L. James Lee, professor of chemical and biomolecular engineering at Ohio State University.
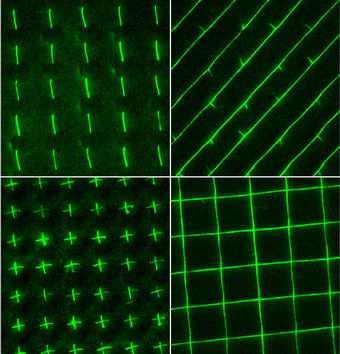
Image above: DNA strands fluoresce in these microscope images from Ohio State University. Researchers here have invented a process for uncoiling DNA strands and forming them into precise patterns – a prelude to biologically based electronics and medical devices. The squares in the lower right image measure approximately 10 micrometers across. Image courtesy of Ohio State University.
In the early online edition of the Proceedings of the National Academy of Sciences, Lee and postdoctoral researcher Jingjiao Guan describe how they used a tiny rubber comb to pull DNA strands from drops of water and stamp them onto glass chips.
Other labs have formed very simple structures with DNA, and those are now used in devices for gene testing and medical diagnostics. But Lee and Guan are the first to coax strands of DNA into structures that are at once so orderly and so complex that they resemble stitches on a quilt.
"These are very narrow, very long wires that can be designed into patterns for molecular electronics or biosensors," Lee said. "And in our case, we want to try to build tools for gene delivery, DNA recombination, and maybe even gene repair, down the road."
The longest strands are one millimeter long, and only one nanometer thick. On a larger scale, positioning such a long, skinny tendril of DNA is like wielding a human hair that is ten meters (30 feet) long. Yet Lee and Guan are able to arrange their DNA strands with nanometer precision, using relatively simple equipment.
In this patent-pending technology, the researchers press the comb into a drop of water containing coils of DNA molecules. Some of the DNA strands fall between the comb's teeth, so that the strands uncoil and stretch out along the surface of the comb as it is pulled from the water.
They then place the comb on a glass chip surface. Depending on how they place the comb, they leave behind strands of different lengths and shapes.
"Basically, we're doing nanotechnology using only a piece of rubber and a tiny droplet of DNA solution," Guan said.
Computer chips that bridge the gap between the electronic and the biological could make detection of certain chemicals easier, and speed disease diagnosis. But first, researchers must develop technologies to mass produce DNA circuits as they produce chip circuits today.
The technique that Lee and Guan used is similar to a relatively inexpensive chip-making technology called soft lithography, where rubber molds press materials into shape.
In this study, they arranged the DNA into rows of "stitches," pinstripes and criss-cross shapes.
The pinstripes presented the researchers with a mystery: for some reason, thorn-like structures emerged along the strands at regular intervals.
"We think the 'thorns' may be used as interconnects between nanowires, or they could connect the nanowires with other electronic components," Guan said. "We are not trying to eliminate them, because we do not think they are defects. We also believe their formation is controllable, because they are almost completely absent in some experiments but abundant in others. Although we currently do not know exactly how the thorns form, maybe new and useful nanostructures may be created if we can better understand and control this process."
The university will license the technology for further development. Lee and Guan are working on their first application – building the wires into sensors for detecting disease biomarkers. In the meantime, they are collaborating with researchers in the Department of Electrical and Computer Engineering at Ohio State to measure the electrical properties of the DNA wires. They are also using this technique to produce DNA-based nanoparticles for gene delivery.
Source: Ohio State University