High Resolution 4Pi Microscopy Reaches the Nucleus
Confocal light microscopy has been an important tool for biomedical scientists as they work to unravel molecular events occurring within human cells. Less than two decades ago, an important advance in microscopy technology was achieved with Dr. Stefan Hell’s invention of “4Pi microscopy.”
Recently developed and commercialized by Leica Microsystems, 4Pi technology offers a significant increase in resolution; however, fundamental optical limitations prevented scientists from visualizing molecules in the cell’s nucleus using this new method.
Now, through the work of Brian Bennett, PhD, of the University of Massachusetts Medical School (UMMS) Department of Biochemistry & Molecular Pharmacology and Leica Microsystems, UMMS Associate Professor of Biochemistry & Molecular Pharmacology Kendall Knight, PhD, and collaborator Jörg Bewersdorf, PhD, of the Institute for Molecular Biophysics at The Jackson Laboratory, this limitation has been overcome—for the first time high resolution 4Pi microscope images of endogenous nuclear proteins in human cells have been realized.
In “H2AX chromatin structures and their response to DNA damage revealed by 4Pi microscopy,” published online this week by the Proceedings of the National Academy of Sciences, Drs. Bewersdorf, Bennett and Knight used the Leica TCS 4Pi microscope to examine histones, nuclear proteins that are involved in packaging and protecting DNA as well as in processes that repair damaged DNA. Exposure to environmental carcinogens can lead to a particularly devastating form of DNA damage, DNA double-strand breaks. The occurrence of such DNA breaks invokes a rapid and highly coordinated series of molecular events within the nucleus, and defects in this process correlate with increased risks for cancer, as well as developmental and immunological abnormalities.
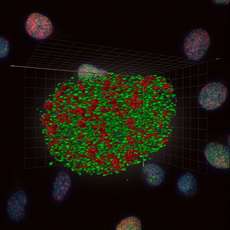
Bewersdorf, Bennett and Knight examined a specific histone protein, H2AX. Within seconds after DNA damage has occurred, H2AX undergoes a subtle but functionally significant molecular modification, which in turn plays an important role in coordinating the recruitment of proteins that will repair the break. Prior to these results, researchers had been unable to visualize how H2AX is distributed throughout the nucleus. However, this study provides never-before-seen high resolution 3D images of H2AX, demonstrating that it exists in distinct clusters that are themselves evenly distributed throughout the entire nuclear volume. This finding suggests that H2AX clusters provide a platform from which signaling and repair events can be coordinated. Additionally, high resolution 4Pi images of modified H2AX reveal for the first time the complex three-dimensional character and dynamics of molecular associations that occur at the site of a DNA break.
“We are very excited to have finally overcome the obstacle that had prevented visualization of nuclear proteins,” said Bennett, who devised the solution enabling the use of 4Pi microscopy during his PhD thesis work in the Knight lab. “The collaborations between myself, Drs. Knight and Bewersdorf, and those at Leica Microsystems were critical in achieving this first step. We look forward to continuing our investigations with the analysis of other nuclear proteins involved in cancer prevention and the repair of DNA damage.”
Source: University of Massachusetts Medical School, Worcester















