Molecular DNA Switch Found to be the Same for All Life
The molecular machinery that starts the process by which a biological cell divides into two identical daughter cells apparently worked so well early on that evolution has conserved it across the eons in all forms of life on Earth. Researchers with the U.S. Department of Energy’s Lawrence Berkeley National Laboratory and the University of California at Berkeley have shown that the core machinery for initiating DNA replication is the same for all three domains of life - Archaea, Bacteria and Eukarya.
In two papers that will be concurrently published in the August edition of the journal Nature Structural and Molecular Biology (now available on-line), the researchers report the identification of a helical substructure within a superfamily of proteins, called AAA+, as the molecular “initiator” of DNA replication in a bacteria, Escherichia coli (E. coli), and in a eukaryote, Drosophila melanogaster, the fruit fly. Taken with earlier research that identified AAA+ proteins at the heart of the DNA replication initiator in archaea organisms, these new findings indicate that DNA replication is an ancient event that evolved millions of years ago, prior to when Archae, Bacteria and Eukarya split into separate domains of life.
“The ability of a cell to replicate its DNA in a timely and faithful manner is fundamental for survival, but, despite decades of study, the structural and molecular basis for initiating DNA replication, and the degree to which these mechanisms have been conserved by evolution have been ill defined and hotly debated,” said biophysicist Eva Nogales, a collaborator on the Drosophila study.
Said biochemist Michael Botchan, also a collaborator on the Drosophila study, “Our two papers fuse together a number of biophysical research techniques to take our understanding of the mechanics of DNA opening and replisome construction to a new level.”
Biochemist and structural biologist James Berger, a participant in both studies added, “Our findings of evolutionary kinship between the DNA initiators in all three domains make sense because, to paraphrase Francois Jacob, the one thing a cell wants to do is to become two cells. A cell can't do this if it doesn't replicate its DNA in the right place, at the right time, and in the right manner, while simultaneously avoiding over-replication.”
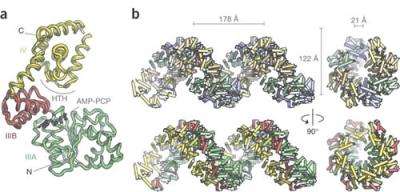
The Drosophila results were reported in a paper entitled: Nucleotide-dependent conformational changes in the DnaA-like core of the origin recognition complex. This study was led by Botchan and Nogales, and included Megan Clarey, Jan Erzberger, Patricia Grob, Andres Leschziner and Berger. Nogales and Berger hold appointments with Berkeley Lab’s Life Science and Physical Biosciences Divisions, respectively, and with UC Berkeley’s Molecular and Cell Biology Department, in which Botchan is a professor. Nogales is also an investigator with the Howard Hughes Medical Institute.
The E.coli results were presented in a paper entitled: Structural basis for ATP-dependent DnaA assembly and replication-origin remodeling. Berger led this study and his collaborators included Erzberger and Melissa Mott.
While the research studies behind these two papers were not coordinated, they did benefit from “a convenient congruence of timely results,” as Berger explained.
“We had solved our initiator structures in the E.coli study just as results were being generated from the Botchan and Nogales groups on the Drosophila study. Once we compared notes, we immediately pooled forces. When we subsequently were able to dock our bacterial model into a region of their eukaryotic structure, it solidified the evolutionary and functional similarities between the two mechanisms.”
For the E.coli study, Berger and his team utilized the exceptionally bright and intense x-rays of Beamline 8.3.1 at Berkeley Lab’s Advanced Light Source synchrotron. With the data gathered at this protein crystallography facility, Berger and his team assembled a high-resolution model of the molecular structure of a protein known as DnaA, which is a member of the AAA+ family. While it has long been known that DnaA controls the process of initiating DNA replication in bacteria, the molecular details of its myriad activities have until now been a mystery.
Berger’s team found that when the DnaA protein binds with adenosine triphosphate or ATP, the nucleotide molecule that supplies energy to all components of a cell, the ring-shaped AAA+ proteins assemble into a right-handed spiraling superstructure. This arrangement was unexpected, because in other functional AAA+ complexes, the ring assemblies are closed. In addition, the architecture indicated that the AAA+ superhelix will wrap coils of the DNA double-helix around its exterior, causing the familiar “spiral staircase” of the DNA to deform as a first step in the separation and unwinding of its two gene-carrying strands.
“It is likely that the AAA+ rings of the replication initiators are open to allow others proteins to dock onto the initiator complex,” said Berger. “These other proteins can help add layers of complexity, such as assisting with helicase loading or inactivating the initiator after replication has begun.The open rings also probably allow DNA to interact with the interior of the initiator assembly.” Bacterial cells, like the cells of Achaeans, are prokaryotes, meaning their DNA is not contained within a defined nucleus. Eukaryotes consist of plants and animals and all other organisms whose DNA is contained within a membrane-bound nucleus. Whereas DNA replication in bacteria is typically initiated at a single sites, DNA replication in eukaryotes can be an immensely complicated multi-event affair, involving the coordinated initiation and regulation of hundreds and even thousands of protein machines at different sites throughout the genome. Furthermore, the highly packaged nature of eukaryotic genomes makes it difficult for these protein machines to access the DNA. Because of this complexity, the mechanism for initiating DNA replication in eukaryotes was presumed to be much different than the prokaryote initiator.
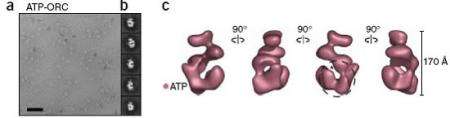
Studies over the past decade have demonstrated that all of the multiple events that initiate DNA replication in a eukaryote are directed by a single complex of proteins called the origin recognition complex (ORC). However, until now, models of the ORC proteins have lacked sufficient detail to identify the structure of the initiator. In their Drosophila study, Nogales and Botchan and their collaborators studied fruit fly ORC using single-particle electron microscopy. Their images revealed for the first time how the ORC when bound to ATP forms a AAA+ helical structure much like the DnaA superhelix found by Berger and his team in their E.coli study.
“This work provides the first view of the mechanical transitions in ORC driven by ATP in a higher organism,” said Nogales. “While our studies have not yet shown the initiator wrapped around the DNA, the structural similarity to the DnaA initiator found in the E.coli study suggests that there are likely to be strong mechanistic commonalities in the ways that initiators engage and remodel replication origins, as well as in how they facilitate replisome assembly.”
The idea that all three domains of life share the same DNA replication initiator is new and will require some re-thinking on the part of biologists who study eukaryotes. Re-thinking will also be required for models of DNA replication that predicted initiators would have similar structures to the protein “clamps” and “clamp loaders” already identified as key mechanisms in the DNA replication process.
Said Berger, “Our work shows that there are major structural distinctions between assembled initiator and clamp loader complexes. This not only has important implications for the respective functions of these different mechanisms, it also calls into question some cherished models in the field.”
The two studies by Nogales, Berger, Botchan and their colleagues also show how when nature finds a mechanism that works well, such a mechanism is conserved through evolution.
Said Nogales, “The specialization of DNA replication initiators took place a long time ago, separating them from other members of the AAA+ superfamily of proteins while maintaining an identity among themselves that reflects the importance of the replication process. Through the millions of years, evolution has added bells and whistles around this highly conserved central engine.”
The E.coli study was supported by the G. Harold and Leila Y. Mathers Charitable Foundation and the National Institutes of Health (NIH). The Drosophila study was also supported by NIH, plus the U.S. Department of Energy’s Office of Biological and Environmental Research and HHMI.
Source: Berkeley Lab

















