Large scale flu virus sequencing completed
Scientists at the Institute for Genomic Research in Maryland report completing the first large-scale project to sequence the influenza virus.
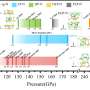
TIGR scientists say they sequenced 209 complete genomes of the human influenza A virus, representing virus samples taken from patients who visited county clinics across New York during five seasons 1999-2004.
Nearly all of the genomes represented the H3N2 strain, which predominated during those flu seasons. Comparing genomes, the researchers tracked the changing virus as it moved across the region.
"This study demonstrates that genomics can help us better track the flu virus and develop more effective vaccines," said first author Elodie Ghedin, who heads TIGR's viral genomics lab. "This is perhaps the most detailed snapshot scientists have gotten of flu's movement through communities."
Across New York state, the researchers documented at least three distinct variants of the H3N2 influenza virus during the 5-year study period. In some of the flu seasons studied, the variants circulated simultaneously. That, the scientists said, means New Yorkers weren't all contracting the same flu but slightly different versions of the virus.
The study appears in the current issue of the journal Nature.
Copyright 2005 by United Press International