What do Biological Cells and Democracy Have in Common?
(PhysOrg.com) -- Scientists at the New England Complex Systems Institute (NECSI) and Harvard University offer a groundbreaking new perspective on how genes determine and regulate the functional identity of a cell. The study, titled Attractors and Democratic Dynamics, appears in the latest issue of Science. It examines views of cell regulation and suggests a model that accounts for a cell's robustness to random perturbations and its sensitivity to specific environmental changes.
The process by which a cell becomes differentiated into, for example, a muscle cell, a neuron, or a blood cell is largely determined by the expression, or "transcription," of genes. This mechanism is of prime importance to many biologists, including developmental biologists studying stem cells and oncologists concerned with how normal cells become cancerous.
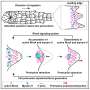
The article shows that to accurately study transcriptional regulation, researchers must consider the interplay between individual genes and collective states. Dr. Yaneer Bar-Yam, one of the article's authors and president of NECSI, explains, “A biological cell is a self-governing community of molecules, like a society. As a society, cell regulation is intermediate between a centrally controlled autocratic government and a democracy. What we need is a way to study cells that take both individual and collective gene action into account.”
Dr. Bar-Yam compares different scenarios to different methods of seeking political change: "Some people who want to make a policy change appeal to those in power--the ministers, the kings, the leaders. Others go to the people and try to affect popular opinion. In reality, both influence the behavior of a nation, and the interactions between individuals and interest groups should be taken into account." Indeed, a cell's robustness to random influence coupled with its ability to quickly respond to certain cues can be explained by the interplay between individual genes and collective states. While the concerted influence of thousands of genes keeps cells steady and protected against random noise, the master genes can cause significant changes so the cell quickly adapts to changing conditions. To incorporate these ideas into current research, the authors describe control coefficients that measure change in collective states relative to individual gene transcription level changes.
There are also important implications for medical research. Dr. Benjamin de Bivort of Harvard University and NECSI, another of the article's authors, believes this more complete picture of cell regulation can have a significant impact on the future of medicine, "With this framework we can better understand how a cell attains, and maintains, its identity. From oncologists to stem cell researchers, scientists can use this framework to create more effective methods for predicting, manipulating, and controlling the fate of cells." Dr. Dion Harmon of NECSI, another of the article’s authors, agrees, "The implications are wide-ranging. These findings could one day help develop better medications that target a large number of genes to cure diseases like cancer."
Provided by New England Complex Systems Institute