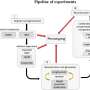
Moss protein plays role in Alzheimer's Disease
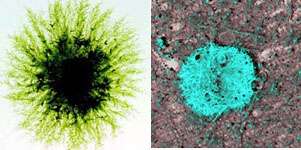
Preventing Alzheimer's disease is a goal of Raphael Kopan, Ph.D., professor of molecular biology and pharmacology at the Washington University School of Medicine. The moss plant Physcomitrella patens studied in the laboratory of Ralph S. Quatrano, Ph.D., the Spencer T. Olin Professor and chair of the biology department on WUSTL's Danforth Campus, might inch Kopan toward that goal. Here's how.
The gene presenilin (PS) in mammals provides the catalytic activity for an enzyme called gamma secretase, which cleaves, or cuts, important proteins Notch, Erb4 and the amyloid precursor protein (APP), all key components of communication channels that cells use to arbitrate functions during development. There are two mammalian genes that occur in mammals for which mutations cause an earlier onset of Alzheimer's. One is APP, where a fragment of the protein accumulates in amyloid plaques associated with the disease. Another common site for mutations is found in PS proteins. The enzyme gamma secretase contains PS and works to dispose of proteins stuck in the cellular membrane.
This enzyme, with PS at its core, mediates two cellular decisions. One is to cut APP and, as a byproduct, generate the bad peptide associated with Alzheimer's; the other is to cut the Notch protein in response to specific stimuli. Notch is then free to enter the nucleus of cells where it partakes in regulating normal gene expression. Without Notch activity, a mammal has no chance of living.
Notch is a part of a short-range mammalian communication channel, and for years it has been known to have a working relationship with PS. However, Notch is absent in plant cells, and PS function in plants remained mysterious until Quatrano's post-doctoral researcher, Abha Khandelwal, Ph.D., arrived at WUSTL and was interested in understanding signal transduction in plants.
"When I searched the literature, the plant signal transduction pathways were not very well documented compared to the mammalian counterparts such as Notch," said Khandelwal. "Meanwhile, my husband, Dilip Chandu, Ph.D., was working in the Kopan lab on ways to study functions of PS without interference from its predominant substrate Notch."
This encouraged Khandelwal to search for the PS gene in the genomes of plants including the recently sequenced Physcomitrella patens genome, to which the Quatrano lab had access. In addition to the known Arabidopsis PS, she found the gene in Physcomitrella and asked, "What is PS doing in moss? Is it acting as an enzyme or does it have a different function?"
Forming a collaboration
"Moss, like yeast, has this great ability where you can actually select a gene and remove it, mutate it, or replace it with another gene from any source. This approach allows us to discern a gene's value and function in moss," said Quatrano, who was a world leader in the sequencing of the moss genome. "It is an excellent system to experimentally discern gene function because of this property as well as others that we and a worldwide consortium have developed over the last several years."
Thus, a collaboration was born. By engaging the expertise of the team in the Kopan lab, the Quatrano lab proceeded to experiment with PS in moss, which finally resulted in a fruitful collaboration recently reported in the Proceedings of the National Academy of Science. Khandelwal proceeded to remove PS, and the result was an obvious change — a phenotype. Moss lacking PS looked different, growing with straight, rigid filaments instead of curved and bent filaments like the parent moss with the PS gene intact.
"That showed the gene has an obvious function that clearly did not require Notch. We just don't know exactly what it is yet, but we have proposed a hypothesis to be tested," Quatrano said.
The phenotype piqued Kopan's interest: He saw the potential of looking at the role of PS independent of Notch. Khandelwal and Chandu took the phenotype, switched out a mammalian form of PS into the phenotype and rescued it. Similarly, inserting the moss gene in mammalian cells resulted in reversing some of the losses experienced by animal cells lacking PS function, testifying that the human and moss proteins had an evolutionary conserved function.
"In the moss, the proteins were very nearly interchangeable," Quatrano said. "This suggested that PS has a role outside the Notch pathway and may provide clues in mammalian systems as to its primary role, independent of its substrate in mammalian cells."
"We were amazed to realize that genes from moss and humans were not only structurally conserved but also shared similar functions," Khandelwal said.
Moonlighting protein in mammals
"We spent a lot of time trying to find an activity of PS to circumvent cleavage of APP, which has been very difficult, "Kopan said. "Importantly, the human protein acted in plant cells even if its enzymatic activity was removed by mutation. We stumbled upon an observation that PS proteins in mammals can perform other functions besides the enzymatic ones, that is, outside its role as gamma secretase. We're now looking closely to define these moonlighting functions and determine their contribution to disease."
In moss, the mutant phenotypes suggest PS might play a role in signal gathering, cytoskeleton organization and/or cell wall composition and organization. Quatrano and Khandelwal are investigating. Kopan, Chandu and others are searching for PS's moonlighting activities in mammalian cells.
"As a developmental biologist, my job is to translate the genetic code as if it were a manufacturer's manual, and that is accomplished by gaining detailed understanding of genes and protein function," Kopan said. "Unfortunately, we're doing it one gene at a time, slowly building networks, figuring out what the context is. We can't think of all of it at once. We have to look at a small subset of genes and how they work with their 'friends', and hope that our observations will fit together in one coherent network."
Quatrano said the collaboration between the two labs is a reflection of what the Genomic Age can do.
"Today, sitting at your computer, you can data mine genomes from hundreds of microorganisms, animals, fungi, insects and plants, and you're seeing more evidence of genes being conserved in widely different organisms," Quatrano said. "This collaboration is a perfect example of bringing two labs together that on the surface have nothing in common other than one protein and two people who were aware of the interests of the other. It's led to a significant contribution that hopefully will lead to further clues as to the function of PS."
With this study, the Kopan and Quatrano labs and others could use this outstanding plant model not only to understand some of the off-target affects during Alzheimer's Disease therapy, but also to unravel novel interactions and pathways in plants.
Source: Washington University School of Medicine