Evolution with a restricted number of genes
The development of higher forms of life would appear to have been influenced by RNA polymerase II. This enzyme transcribes the information coded by genes from DNA into messenger-RNA (mRNA), which in turn is the basis for the production of proteins. RNA polymerase II is highly conserved through evolution, with many of its structural characteristics being conserved between bacteria and humans.
Single-cell organisms were already in existence 500 million years ago, with several thousand genes providing different cellular functions. Further developments seemed dependent on producing even more genes. For a highly developed organism like a human, this form of evolution would have resulted in several million genes.
Researchers were therefore surprised to learn, following publication of the human genome, that a human only has around 25,000 genes – not many more than a fruit fly or a worm with approximately 15,000 to 20,000 genes. It would appear that, over the last 500 million years, other ways to produce highly complex organisms have evolved. Evolution has simply found more efficient ways to use the genes already there. But what could have made this possible?
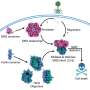
In the current issue of Science the group of Prof. Dirk Eick at the Institute of Clinical Molecular Biology and Tumor Genetics, GSF – National Research Center for Environment and Health, Munich, and the group of Dr. Shona Murphy from Oxford University, England, publish results which represent a piece of the puzzle and shed new light on to the purpose of an unusual structure in RNA polymerase II. They build on earlier observations that gene expression is not just regulated by binding of the enzyme to the gene locus to which it is recruited, but also during the phase of active transcription from DNA into RNA.
During this phase, parts of the newly synthesised RNA may be removed and the remaining sequences combined into new RNA message. This ‘splicing’ of RNA occurs during gene transcription, and in extreme cases, can produce RNAs coding for several thousand different proteins from a single gene.
But what was the development that permitted this advance in gene usage? The RNA polymerase II has developed a structure composed of repeats of a 7 amino-acid sequence. In humans this structure – termed “carboxyterminal domain” or CTD – is composed of 52 such repeats. It is placed exactly at the position where RNA emerges from RNA polymerase II. In less complex organisms the CTD is much shorter: a worm has 36 repeats, and yeast as few as 26, but many single-cell organisms and bacteria have never developed an obvious CTD structure.
Although the requirement of CTD for the expression of cellular genes in higher organisms is undisputed, the molecular details for the gene-specific maturation of RNAs is still largely enigmatic. The groups of Dirk Eick and Shona Murphy have now shown a differential requirement for phosphorylation of the amino acid serine at position 7 of CTD in the processing and maturation of specific gene products.
These results provide the groundwork for the discovery of further pieces of the CTD puzzle and thus enlarge our knowledge of gene regulation. Given its fundamental importance, understanding the mechanism of gene regulation is essential if we are to understand cancer and other diseases at the molecular level and develop new therapies.
Source: GSF - National Research Center for Environment and Health