New technique promises to aid doctor's ability to identify, treat bacterial infections
A new technique developed by a University of Central Florida chemist will help physicians more quickly identify the bacterial infections patients have so they can be treated in hours instead of days.
As more bacterial strains resistant to many drugs emerge, it becomes more critical to quickly identify infections and the antibiotics that would most effectively treat them. Such quick identifications become even more important during epidemics because large numbers of samples would have to be tested at once.
Assistant Professor J. Manuel Perez’s new technique also promises to give research institutes and pharmaceutical companies a quicker and cheaper way of developing new antibiotics to combat super bugs.
The results of Perez’s study were recently published online in Analytical Chemistry (http://pubs.acs.org/cgi-bin/asap.cgi/ancham/asap/pdf/ac701969u.pdf). The research was funded in part by the National Institutes of Health.
“The method really gives doctors quicker access to test results so they can treat their patients more quickly,” Perez said from his lab at the Nanoscience Technology Center at UCF. “But there are more applications. This method can also be used by research facilities and big pharmaceutical companies for the high throughput screening of drugs for antibacterial activity.”
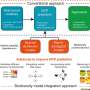
Perez uses gold nanoparticles coated with a sugar and a protein that binds to sugars. Meanwhile, a variety of antibiotics are placed in the same solution. A spectrophotometer reads optical variations in the gold nanoparticle solution as the sugar and protein shift , which in turn demonstrate which antibiotics effectively halt bacteria growth and which ones do not. Results can be obtained within a couple of hours, in contrast to the traditional methods, which can take days to complete. And hundreds of samples can be tested at once using this technique because the amount of bacteria and antibiotic needed is small.
Pharmaceutical companies can use existing equipment to read the variations, which means they do not have to buy new equipment. Perez’s study also shows that the technique is as sensitive and accurate as the traditional, more time-consuming approach.
“We’re very excited and very pleased with the results,” Perez said.
Source: University of Central Florida