Understanding why C. difficile causes disease -- it's Hungry
Researchers studying the genetics behind why C. difficile causes disease have come to a simple conclusion -- the bacteria do it because they are starving. That just might help them find a new treatment for what can sometimes be a very difficult disease to treat.
"The genes responsible for toxin production only seem to be expressed during periods of nutrient deprivation. This is consistent with the view that most disease-causing bacteria express their pathogenicity when they are hungry," says Abraham Sonenshein, professor at the Sackler School of Graduate Biomedical Sciences at Tufts University and at Tufts University School of Medicine, at the 107th General Meeting of the American Society for Microbiology (ASM) on May 24, 2007.
C. difficile bacteria are everywhere — in soil, air, water, human and animal feces, and on most surfaces in hospital wards. The bacteria don't cause problems until they grow in abnormally large numbers in the intestinal tract. This can happen when the benign bacteria that normally inhabit the intestinal tract are reduced such as when people take antibiotics or other antimicrobial drugs. Then, C. difficile can cause symptoms ranging from diarrhea to life-threatening inflammations of the colon.
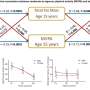
In 2002 a new, more virulent strain began appearing in hospitals in the United States and Canada. Recently, this strain was shown to be responsible for more than half of all cases in a representative sampling in Quebec. The highly virulent strain has a much higher toxin production which leads to more destructive and deadly disease, says Vivian Loo of McGill University.
Sonenshein is studying a five-gene region of the bacterium’s chromosome known as the tcd locus. Two of the genes code for the toxins the bacterium produces that cause disease and a third gene codes for a protein that makes a hole in the organism’s cell envelope to let the toxins out. The last two genes are of greatest interest to Sonenshein and his colleague, Bruno Dupuy from the Institut Pasteur. One codes for a protein, known as R, that is necessary for the expression of the first three genes and the other codes for a protein called C that prevents R from acting.
A mutation in the C protein gene, leaving R unchecked, is the cause of the hypervirulent strain. Sonenshein and his colleagues are currently working to identify a protein that might shut down the gene that codes for R. By identifying such a protein, Sonenshein hope to find a way to change the appetite of the bacteria. "If we find a way to shut down toxin production in the hypervirulent strain, we might have a new way to treat the disease," says Sonenshein.
Source: American Society for Microbiology