Gene hunters close in on Lou Gehrig’s disease
In the first genome-wide search for the genetic roots of the most common form of amyotrophic lateral sclerosis (ALS), Johns Hopkins scientists have newly identified 34 unique variations in the human genetic code among 276 unrelated subjects with ALS.
The 34 so-called single nucleotide polymorphisms, or SNPs, represent good candidate genes predisposing people to the non-inherited form of the fatal neurodegenerative disease.
“Although we haven’t located the exact gene responsible for sporadic ALS, our results seriously narrow the search and bring us that much closer to finding what we need to start developing treatments for the disease,” says Bryan J. Traynor, M.D., of the Department of Neurology at The Johns Hopkins University School of Medicine.
ALS, also known as Lou Gehrig’s disease for the legendary Yankee first baseman who succumbed to its paralyzing assault on the nervous system, kills 10,000 Americans a year. An estimated one in 2,000 people is at risk of developing the disease.
Genes behind inherited forms of ALS -- responsible for about only 5 percent of all cases -- were discovered a decade ago, but no genetic roots have previously been found for sporadic ALS, which occurs in people without a family history of the disease.
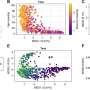
In the Johns Hopkins study, described in the online version of Lancet Neurology this month, Traynor and his team scanned the entire genome of 276 adult male and female subjects with sporadic ALS and 271 adult male and female subjects with no history of neurological disease.
Researchers used a new technology known as “SNP chips” to analyze all 555,352 SNPs in the genome of each subject. SNP chips are glass slides that resemble computer processor chips. They’re coated with tiny beads that “read” the SNPs scattered throughout the human genome.
The researchers found 34 genetic variants that ALS patients were more likely to have compared to normal individuals without the disease.
“This is the first major step toward understanding how genetics may influence the most common form of ALS,” says co-researcher Jeffrey D. Rothstein, M.D., Ph.D., professor in the Department of Neurology at Johns Hopkins. “The results will not only help us to better understand sporadic ALS but also to tailor tools that will reveal therapies for it.”
Traynor cautioned that the 34 SNPs are not guaranteed trail markers for ALS genes. “If you roll dice 555,352 times, you are bound to get lucky by chance alone on some of those throws,” he says. “The next step is to go back and figure out which of these ‘hits’ are real and which are false.”
Thousands of other SNPs were more weakly associated with ALS, some of which could turn out to be just as important as those more strongly linked.
Traynor says a follow-up study is in the works that will replicate the research using a similar number of patients and controls. Traynor, who has a joint affiliation with the National Institute of Mental Health in Bethesda, Md., collaborated in this study with lead author Jennifer Schymick and fellow researcher John Hardy, Ph.D., both of the Laboratory of Neurogenetics at the National Institute on Aging in Bethesda. Researchers from the National Institutes of Health, from the Packard Center for ALS Research at Johns Hopkins and from the University of Turin, Italy, also contributed to this study.
Source: Johns Hopkins Medical Institutions