UGA researchers use patented SERS technique to rapidly, accurately detect rotavirus strain
Using nanotechnology and a patented signal enhancing technique developed at the University of Georgia, UGA researchers have discovered a rapid, sensitive and cost-effective method to detect and identify a number of rotavirus strains and genotypes in less than one minute with greater than 96 percent accuracy.
In their study, Ralph A. Tripp and Jeremy D. Driskell, researchers in the College of Veterinary Medicine's department of infectious diseases, and Yiping Zhao and Richard Dluhy, researchers in the Franklin College of Arts and Sciences departments of physics and chemistry, utilized surface enhanced Raman scattering, or SERS, to detect and quantify Group A rotaviruses.
Group A rotaviruses are the leading cause of severe gastroenteritis in infants and young children, infecting approximately 130 million children annually. Rotavirus infections are responsible for approximately 2 million hospitalizations and more than 500,000 deaths each year, and are particularly burdensome on health care resources in developing countries. Clinical diagnostic tests currently used to detect rotavirus do not provide information on the genotypes, which is essential for aiding public health officials in monitoring epidemics, identifying novel strains and controlling disease.
Tripp and Driskell worked with the most commonly identified strains of rotavirus, provided by Carl D. Kirkwood of the Murdoch Childrens Research Institute, at the Royal Children's Hospital in Parkville, Australia, to show that SERS can detect and identify numerous virus strains and genotypes in less than 30 seconds, without the need to amplify the analyte for detection. Their technique requires no or minimal specimen preparation for analysis and uses minimal volumes of analyte.
"Nanotechnology has provided a considerable advance in diagnostic and prognostic capabilities," noted Tripp. "The technology strengthens and expands current diagnostic applications by providing a means to enhance existing technology for novel applications such as SERS detection of viruses. The field of diagnostics and biosensing has been pushed dramatically forward by our ability to now amplify and detect the molecular fingerprints of pathogens as opposed to amplifying the pathogens for detection."
The findings from the UGA research team are important as most enteric viruses produce diseases that are not readily distinct from other pathogens and diagnostics are generally limited to attempts at viral culture, antibody-mediated antigen detection and polymerase chain reaction. These methods are cumbersome, often have limited breadth and sensitivity in detection and/or offer limited information on genotype.
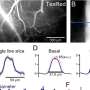
SERS works by measuring the change in frequency of a near-infrared laser as it scatters off viral nucleic acid and protein components. This change in frequency, named the Raman shift for the scientist who discovered it in 1928, is as distinct as a fingerprint.
More information: The study was published in PLoS ONE on April 19.