Top notch decisions in the developing airways bring insights into lung disease
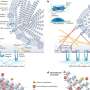
In the normal lung, the airways are lined by a balanced mixture of ciliated, secretory and neuroendocrine cells which perform functions as diverse as air humidification, detoxification, and clearance of environmental particles. This balance can be altered dramatically by faulty adaptation responses of the lung to cigarette smoke or allergens in patients with Chronic Obstructive Pulmonary Disease (COPD) and asthma.
How these different cell types emerge from lung progenitor cells and how these fates are balanced in developing airways, remain an open question. A study from a research team led by Wellington Cardoso, MD, a professor at the Pulmonary Center Boston University School of Medicine and Director of the Program in Lung Development and Progenitor Cell Biology, sheds light into this problem.
The Notch pathway is a major regulator of cell fate decisions in developing cells from fruit flies to humans. Using mouse genetic models, the BU researchers inactivated Notch signaling in the lung and discovered that airways no longer formed secretory cells. Instead they became populated almost exclusively by ciliated cells. The researchers showed that this imbalance seems to result from the loss of a mechanism of cell fate choice triggered by the Notch called lateral inhibition.
"When you lose Notch signaling, you lose the ability to generate secretory cells that make the lining fluid critical for protection and integrity of airway, and the other fate, of ciliated cells is de-repressed" said Dr. Cardoso.
These findings help to understand how airways form and provide insights into how interfering with Notch signaling may be potentially useful as a therapeutic intervention in respiratory diseases, such as asthma and COPD, in which airways have an overabundance of secretory cells and paucity of ciliated cells in the airways. The link between hyperactive Notch and excessive secretion is now rapidly emerging from other recent reports.
Source: Boston University Medical Center