Nine new X chromosome genes associated with learning disabilities
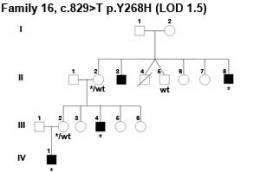
(PhysOrg.com) -- A collaboration between more than 70 researchers across the globe has uncovered nine new genes on the X chromosome that, when knocked-out, lead to learning disabilities. The international team studied almost all X chromosome genes in 208 families with learning disabilities - the largest screen of this type ever reported.
Remarkably, the team also found that approximately 1-2% of X chromosome genes, when knocked-out, have no apparent effect on an individual's ability to function in the ordinary world. The publication in Nature Genetics - a culmination of five years of scientific collaboration - emphasises the power of sequencing approaches to identify novel genes of clinical importance, but also highlights the challenges researchers face when carrying out this kind of study.
Estimates suggest that the prevalence of learning disability is 2-3%. Learning disability is significantly more common in males than in females and genetic causes have long been sought on the X chromosome: males have only one X chromosome and so a gene mutation on the X is more likely to have an effect in males than in females.
"We sequenced 720 out of the approximately 800 known genes on the X chromosome in more than 200 families affected by X-linked learning disabilities," explains Professor Mike Stratton, from the Wellcome Trust Sanger Institute. "This is the largest sequencing study of complex disease ever reported."
In part because of their apparent effects in males, X-linked disorders have been well studied by geneticists over the past 25 years. Conceived by Dr Lucy Raymond, from the University of Cambridge, and Professor Mike Stratton, the collaborative study harnessed DNA sequencing to detect as many new abnormal genes as possible. In the future similar strategy will be used to find disease causing sequence variants implicated in other complex genetic diseases.
Some characteristics in common could be identified in patients with variants in the same gene. However, in many, learning disability was the only feature. The newly identified genes play roles in a wide range of biological processes suggesting that disruption to many cellular machines can damage the nervous system.
"As well as these important new gene discoveries relating to learning disability, we have also uncovered a small proportion - 1% or more - of X chromosome protein-coding genes that, when knocked out, appear to have no effect on the characteristics of the individual," explains Mike Stratton. "It is remarkable that so many protein-coding genes can be lost without any apparent effect on an individual's normal existence - this is a surprising result and further research will be necessary in this area."
This finding will also act as a warning to geneticists. Large-scale studies are designed to uncover associations between knocked-out genes and disease. However, this study shows that a small proportion of gene knock-outs have no discernable effect on the individual. In future studies, researchers must therefore be cautious about assuming that the presence of a knocked out gene in an individual with a particular disease means that the knocked out gene is causing the disease.
The research comes towards the end of a long process of gene cataloguing in this field. Scientists believe that there are likely to be more undiscovered genes that contribute to X-linked learning disabilities; however, variants at what are expected to be lower frequency will become increasingly difficult and costly to uncover. The next challenge is to implement this improved knowledge of the complex of genes that lead to learning disabilities in clinical practice.
"We already offer genetic counselling to families with X-linked learning disabilities," says Dr Lucy Raymond, Reader in Neurogenetics, Cambridge Institute for Medical Research at the University of Cambridge. "This new research uncovers yet more genes that can be incorporated to improve the provision of diagnostics to families with learning disabilities and allow us to develop more comprehensive genetic counselling in the future, allowing parents and the extended family to make the most informed family planning decisions."
Although in most cases improved treatments remain to be developed, information about a genetic condition can provide support to affected families. The information can also help to inform reproductive choices.
More information: Tarpey PS et al. (2009) A systematic, large-scale resequencing screen of the X chromosome coding exons in mental retardation. Nature Genetics. Published online before print as doi: 10.1038/ng.367
Source: Wellcome Trust Sanger Institute (news : web)