By imaging live cells, researchers show how hepatitis C replicates
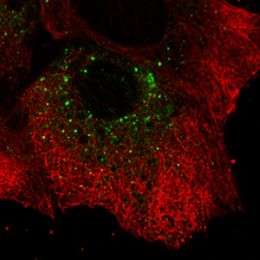
(PhysOrg.com) -- The hepatitis C virus is a prolific replicator, able to produce up to a trillion particles per day in an infected person by hijacking liver cells in which to build up its viral replication machinery. Now new research — in which scientists have for the first time used fluorescent proteins to image hepatitis C virus replication in live cells — shows that the microscopic viral factories are a diverse mix of big, immobile structures and tiny replication complexes that zip zanily around inside the cell. The scientists say their results, published in the Journal of Virology, offer new insights into how this difficult-to-treat virus, and perhaps others in its class, ensures efficient reproduction of itself — knowledge that could help design next-generation treatments.
“There is so much that we don’t know about this virus, so a better understanding of how the pathogen cleverly forms lots of large and small factories to reproduce itself so that it can infect new hosts may be of great benefit,” says study coauthor Benno Wölk, a former postdoctoral researcher in the Rockefeller University Laboratory of Virology and Infectious Disease. He is now a researcher and physician at Hannover Medical School in Germany.
An estimated 170 million people worldwide are chronically infected with hepatitis C, which is a major cause of liver cirrhosis and liver cancer. So progress in understanding and treating the infection is crucial, says the study’s senior investigator, Charles M. Rice, Maurice R. and Corinne P. Greenberg Professor in Virology and director of the Center for the Study of Hepatitis C. “There is no vaccine available for hepatitis C, and current therapies are not always effective because the virus fights back against drugs developed to block replication,” Rice says.
Scientists had until now believed that the virus’s replication process occurred in one or several large complexes inside a cell. It was hard to learn more because in order to see the virus it had to be killed. “Up to this study, researchers have only been able to look at infected cells when they were fixed and immobile,” Wölk says. “They found areas where the cell membrane was altered and found viral proteins in these structures that suggested that was where replication took place.”
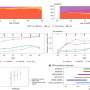
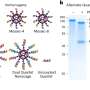
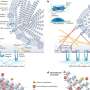
To visualize the replication process, the researchers selected one of the proteins that the hepatitis C virus uses to make its replication factories and fused it to a green fluorescent protein, which emits a green glow when exposed to a specific wavelength of light. They were surprised to see that the small hepatitis C virus replication complexes were transported around the cell. “It’s remarkable that the virus hijacks the cell’s transport machinery to move the viral replication complex around,” Wölk says. “We also learned that for the first several hours after infection only small structures, like dots, formed, which were quickly spread all over the cell. Then the big structures took shape, and they didn’t move.”
The researchers theorize that the small structures are the actual sites of viral replication and that the big structures are clusters of the smaller factories — perhaps formed after the virus has already successfully settled in with the cell. “It is questionable whether the virus even needs the big structures to replicate. They could be performing other functions or they could just represent garbage cans of the cell,” Wölk says. “This is very different from the traditional view.”
Although they can’t say for sure, the group, which also includes Benjamin Büchele of the University of Freiburg in Germany, and Darius Moradpour of the University of Lausanne in Switzerland, suspects that these small, mobile, replication complexes are more efficient and elegant than large structures because they do two things: distribute the factories so that the integrity of the cell is maintained, and keep the complexity of the replication factories to independent, small, manageable units that are easier to control for the virus.
What the researchers discovered in the hepatitis C virus may also prove to be true for related single-strand RNA viruses in the Flaviviridae family, Wölk says. “If that is the case, then we may be able to find a new treatment target for not just one, but many viral infections.”
Citation: Journal of Virology 82(21): 10519–10531 (November 2008)
Provided by Rockefeller University