Scientists show why cells starved of iron burn more glucose
Duke University Medical Center scientists have found a mechanism that allows cells starved of iron to shut down energy-making processes that depend on iron and use a less efficient pathway involving glucose. This metabolic reshuffling mechanism, found in yeast cells, helps explain how humans respond to iron deficiency, and may help with diabetes research as well.
"If we can understand what metabolic changes happen along a gradient of iron deficiency, then we might be able to identify signatures of a modest iron deficiency in humans," said lead researcher Dennis J. Thiele, Ph.D., who is the George Barth Geller Professor of the Duke Department of Pharmacology and Cancer Biology, "We could head it off at the pass."
"This basic science discovery in yeast sheds important new light on how humans may respond to iron deficiency, which is the most common nutritional disorder," said Duke School of Medicine Dean Nancy C. Andrews, an expert in human diseases of iron metabolism.
The findings, published in the June issue of Cell Metabolism, are also potentially important for those studying diabetes. "Evidence is growing that if there is an iron imbalance in the beta cells of the pancreas, these cells won't produce insulin properly," Thiele said. "Now we know what happens in yeast in terms of glucose (sugar) utilization. We need to learn whether the same cause and effect holds true in mammals."
Iron deficiency anemia affects nearly 2 billion people worldwide, most often pregnant women, premature babies, and young children, Thiele said. Anemia profoundly affects cognitive development, and motor and neuronal development, he said.
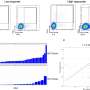
The scientists wanted to know how organisms establish a balance of iron in their cells. "We now know when yeast cells encounter iron deficiency, they reorganize their metabolism by degrading specific messenger RNAs (mRNAs) and leaving other messenger RNAs alone, which begins a sequence of events," said Thiele. Messenger RNAs are molecules that carry coding information from the DNA to the structures that make proteins, which in turn regulate the body's structures and functions.
The first response to iron deficiency is to shut down the energy hub of the cell, the mitochondria, which takes glucose and turns it efficiently into cell energy fuel, or ATP. The mitochondria depend greatly on iron. As a cell becomes more starved for iron, it "dials down" the mitochondrial processes by degrading the mRNAs encoding the proteins involved in such processes, and thus, some iron is freed up, Thiele said.
The second response is to shut down iron storage pathways and other, more dispensable biochemical reactions that depend on iron. "When you are low on iron, you don't want to save it and take it out of use," Thiele explained.
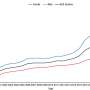
The third response is to increase glucose utilization pathways outside of the mitochondria, which is a much less efficient way to produce energy. Glucose molecules processed for energy outside of the mitochondria create about 18 times less energy, said co-author Sandra Vergara, a doctoral student in Thiele's lab.
"Cellular iron balance follows the rules of economics," Vergara said. "During scarcity, the cell prioritizes the utilization of iron, saving it for more essential processes. This prioritization comes at a cellular cost, which is reflected in the higher demand for glucose, so the cell can keep the correct amount of energy flowing."
If we run low on ATP, we become tired and lethargic, which are symptoms of iron deficiency, Thiele said. "Iron is hard for humans to get from plant sources, which form the basis for most of the world's diet." Iron is very abundant in nature, but cells have a hard time taking it up, because it can change its form inside the body.
Thiele stressed that the findings show what happens during iron deficiency in baker's yeast cells, but probably in some way do extend to people. "Nearly 35 percent of all known human disease genes have a counterpart in the yeast genome. A scientist is always conservative about extrapolating. I think we can make predictions that the metabolic reshuffling that we observe in yeast, the same types of key proteins and enzymes that are involved during iron deficiency, are likely to follow similar patterns in human cells."
Most of the primary metabolism pathways are conserved at the molecular level from yeast to humans, Vergara said.
Source: Duke University