Researchers discover possible markers for mental illness
Researchers have discovered natural genetic differences that might help predict the most effective antipsychotic drugs for particular patients with mental disorders such as schizophrenia, Parkinson’s and drug addiction.
They found the differences in the gene for a molecule called the dopamine D2 receptor (DRD2), a protein present on brain cells that are sensitive to the neurotransmitter dopamine.
The receptor is known to play a key role in memory and in a variety of mental illnesses. Most antipsychotic drugs work at least in part by blocking this protein, but scientists don’t yet understand how this helps patients. Nor can they explain why some people respond well to certain antipsychotic drugs and others respond poorly.
“Our study shows that these differences affect normal brain activity and memory processing, and therefore may also be important in mental illness,” says principal investigator Wolfgang Sadee, program director in pharmacogenomics at the Ohio State University Medical Center.
The findings could lead to tests that will enable doctors to match patients with certain mental illnesses to the most effective therapy, something they cannot do now.
The study was done in collaboration with Professor Alessandro Bertolino, University of Bari, Italy, who performed the clinical research. It is published online in the Proceedings of the National Academy of Sciences.
“Identifying these predictive markers is important because antipsychotic drugs are effective in only a portion of patients upon first treatment, and it takes a month or more to establish their efficacy,” says Sadee, who is also a professor of psychiatry and chair of the department of pharmacology.
“During this time, irreparable damage can result if the wrong antipsychotic is given to a patient.”
Sadee notes that the D2 receptor gene has been implicated in mental illness for some time, but that a variety of clinical studies have failed to consistently link variations in the gene to disease.
These findings may change that.
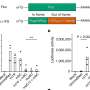
For this study, Sadee and his colleagues analyzed 68 autopsy samples of normal human brain tissue. For each case, the researchers measured and compared the amount of messenger RNA made by each of the two copies of the DRD2 gene. Messenger RNA is a molecule made when a gene is involved in making its protein.
In 15 of the 68 cases, the relative amounts of messenger RNA made by one gene in the pair was strikingly different from the amount made by the other. The disparity was a clue that something was different between the genes.
Comparisons of these DRD2 genes to the rest revealed three small differences in the DNA called single-nucleotide polymorphisms, or SNPs (pronounced ‘snips’).
SNPs are tiny natural variations between individuals that occur at certain positions in genes, providing landmarks in the genome.
SNPs often have no effect on the function of the gene or its protein, but, in this case, laboratory experiments showed that particular changes in two of the SNPs alters how the messenger RNA for DRD2 is processed.
That, in turn, changed the relative amounts of two variants of the protein that are made by the gene.
“The two variants of DRD2 have distinct functions, facilitating or inhibiting dopaminergic transmission, so that a change in their ratios is potentially critical,” Sadee says. “We believed that this change would enhance dopamine activity in the brain.”
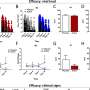
The researchers then tested this hypothesis in normal human volunteers who took simple memory performance tests. The participants’ brain activity was monitored during the testing by functional magnetic resonance imaging (fMRI).
The results showed that volunteers with the two variant SNPs had significantly more brain activity than the usual SNPs for the same memory task.
“Their brain needed to ‘work’ more to get the same result,” Sadee says. The two SNPs were also associated with reduced memory performance and attentional control.
Sadee and his colleagues are now testing the relevance of the SNP markers in patients with schizophrenia and in patients with cocaine addiction.
Source: Ohio State University