One man's junk may be a genomic treasure
Scientists have only recently begun to speculate that what’s referred to as “junk” DNA – the 96 percent of the human genome that doesn’t encode for proteins and previously seemed to have no useful purpose – is present in the genome for an important reason. But it wasn’t clear what the reason was. Now, researchers at the University of California, San Diego (UCSD) School of Medicine have discovered one important function of so-called junk DNA.
Genes, which make up about four percent of the genome, encode for proteins, “the building blocks of life.” An international collaboration of scientists led by Michael G. Rosenfeld, M.D., Howard Hughes Medical Investigator and UCSD professor of medicine, found that some of the remaining 96 percent of genomic material might be important in the formation of boundaries that help properly organize these building blocks. Their work will be published in the July 13 issue of the journal Science.
“Some of the ‘junk’ DNA might be considered ‘punctuation marks’ – commas and periods that help make sense of the coding portion of the genome,” said first author Victoria Lunyak, Ph.D., assistant research scientist at UCSD.
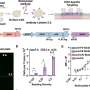
In mice, as in humans, only about 4 percent of the genome encodes for protein function; the remainder, or “junk” DNA, represents repetitive and non-coding sequences. The research team studied a repeated genomic sequence called SINE B2, which is located on the growth hormone gene locus, the gene related to the aging process and longevity. The scientists were surprised to find that SINE B2 sequence is critical to formation of the functional domain boundaries for this locus.
Functional domains are stretches of DNA within the genome that contain all the regulatory signals and other information necessary to activate or repress a particular gene. Each domain is an entity unto itself that is defined, or bracketed, by a boundary, much as words in a sentence are bracketed by punctuation marks. The researchers’ data suggest that repeated genomic sequences might be a widely used strategy used in mammals to organize functional domains.
“Without boundary elements, the coding portion of the genome is like a long, run-on sequence of words without punctuation,” said Rosenfeld.
Decoding the information written in “junk” DNA could open new areas of medical research, particularly in the area of gene therapy. Scientists may find that transferring encoding genes into a patient, without also transferring the surrounding genomic sequences which give structure or meaning to these genes, would render gene therapy ineffective.
Source: University of California - San Diego