Finding of long-sought drug target structure may expedite drug discovery
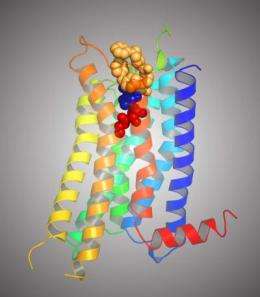
Researchers have solved the three-dimensional structure of a key biological receptor. The finding has the potential to speed drug discovery in many areas, from arthritis to respiratory disorders to wound healing, because it enables chemists to better examine and design molecules for use in experimental drugs.
The researchers are from the National Institutes of Health, collaborating with labs at The Scripps Research Institute and the University of California, San Diego. The finding is published in the March 10 edition of Science Express.
"This is an important step forward — it was impossible until recently to know how this type of receptor is switched on by chemical signals like a tiny machine," said Dr. Kenneth A. Jacobson, chief of the Laboratory of Bioorganic Chemistry in NIH's National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) and an author on the paper. "The architecture of the activated receptor allows us to think in more detailed terms about the other half of the drug interaction. We hope that we're on the verge of a revolution that will expedite the process of crafting new drugs to treat disease."
With this finding, scientists in Jacobson's lab, including co-author Dr. Zhan-Guo Gao, will next work on testing this drug-engineering approach with similar molecules they have newly synthesized.
Jacobson and Gao are part of the NIDDK's intramural program, which enables basic scientists and clinicians of diverse skills and expertise to collaborate on solutions to some of the most difficult issues of human health. Several compounds from Jacobson's lab are currently in clinical trials as potential treatments for conditions including chronic hepatitis C, psoriasis and rheumatoid arthritis.
"Discoveries like this, with the potential to lead to future treatments in a wide variety of areas, are why NIH funds basic science," said NIDDK Director Dr. Griffin P. Rodgers. "By understanding the body at its smallest components, we can learn how to improve whole-body health."
A receptor is a protein that receives and sends signals to other molecules. The three-dimensional structure of the solved receptor also contains an agonist — a chemical command signal from outside the cell — in this case, an adenosine molecule. Similar to the function of a telephone receiver, the receptor acts as a sensor, picking up the message from the agonist and transmitting its information, which begins processes inside the cell.
The researchers discovered that a previously known agonist molecule would bind to its receptor target in a way that stabilizes the protein for crystallization. Once crystallized, the structure can be seen by bombarding it with X-rays. The agonist solidifies the protein by connecting to multiple parts of the receptor with its molecular arms, in the process initiating the function of the entire structure. This adenosine receptor, called A2A, counteracts inflammation and responds to organs in distress. It belongs to the G-protein coupled receptor family, which is involved in processes necessary for many drugs currently in use to take effect. These findings may lead to new drugs for many diseases.
The research was also supported by the National Cancer Institute and the National Institute of General Medical Sciences, both components of the NIH.
"Long-term NIH technology investments in structural biology, including the Protein Structure Initiative, have brought diverse teams of investigators together and yielded powerful methods like the ones used in this study," said NIGMS Director Dr. Jeremy M. Berg. "Receptors must undergo substantial changes in shape in order to function, and revealing these molecular dances in such great detail is an impressive accomplishment."
















