Embracing our differences
While it may have been a momentous occasion in scientific history, the assembly of the first human genome sequence in 2003 was only a first step toward understanding the extent and biological importance of human genetic variation.
In fact, this ‘reference genome’—also known as NCBI36—was not derived from a single individual, but is instead a patchwork constructed from several anonymous donors. In subsequent years, research groups have taken advantage of increasingly powerful and affordable gene sequencing technology to construct full genomes from several individuals of European, African and Asian ancestry. However, such analyses still face major obstacles, even with the benefit of contemporary technology.
“Although ‘next generation’ sequencers can now sequence a human genome within a couple of weeks, sequencing errors are problematic because they are relatively frequent,” explains Tatsuhiko Tsunoda of the RIKEN Center for Genomic Medicine in Yokohama. “Sophisticated methodologies are necessary for detecting genetic variations, including single-nucleotide, copy number and structural variations.” His concern about these issues is particularly strong given his group’s involvement in the International Cancer Genome Consortium (ICGC), an organization focused on understanding how specific genomic alterations might potentially contribute to the tumor formation and progression.
In partnership with RIKEN colleague Akihiro Fujimoto, Tsunoda developed more sophisticated methods for sequence data analysis. As a test of the effectiveness of their approach, they have now assembled the first complete genome sequence from an individual of Japanese ancestry.
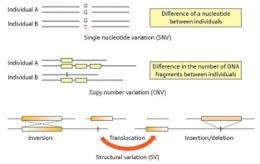
Beyond its status as a landmark in genomics research, this study has also revealed a surprising number of potentially medically relevant sequence and structural variations, both large and small, which have not been identified in previously assembled human sequences. In fact, their analysis of individual NA18943 revealed a striking amount of variability relative to NCBI36. “We found a roughly 0.1% difference between our assembled DNA sequences compared to the reference genome, with approximately three million base-pairs of novel sequences, as well as 3.13 million single-nucleotide variations (SNVs),” says Tsunoda.
Novel SNVs pose a particular challenge to identify, as it is often difficult to be certain whether a putative base change represents a true difference from the reference sequence or is merely the result of an error in the sequencing process. To maximize their accuracy, the researchers carefully compared three different approaches for deciding which base actually occurs at a given genomic position, developing a method that ultimately allowed them to achieve a low rate of false-positive SNV identification.
Notably, a large percentage of the novel SNVs detected in this study represented variations to genes that either disrupt protein production (nonsense mutations) or markedly alter the encoded protein sequence (nonsynonymous SNVs). The researchers hypothesize that such variations are likely to be rare within populations because of their potential contribution to human disease and as such would be strongly selected against over the course of evolution.
Tsunoda and colleagues observed a similar pattern when they compared NA18943 to six other previously characterized individual genomes. Of the nonsense SNVs identified within this collected dataset, 63% were ‘singletons’, or variants that occurred only once across all seven genome sequences. Further, the total collection of nonsynonymous SNVs contained significantly more singletons than were found among the set of non-protein-altering, synonymous SNVs.
Their analysis also revealed numerous regions where the NA18943 genome had been subject to insertions or deletions, more than 350 of which were predicted to markedly alter or disrupt the coding sequence of a gene. Notably, a significant percentage of these were detected within genes involved in olfactory or chemical stimulus perceptions, both of which are known to vary extensively between individuals.
The researchers used a variety of established molecular biology techniques to verify the quality of these data from NA18943. Their findings collectively confirm that the genome of any given individual is likely to exhibit large numbers of rare, but functionally meaningful, variations relative to the general population or even individuals who are closely related from an evolutionary perspective. “We will have to sequence many more individuals within our population as well as across other populations around the world in order to obtain a clearer, more complete picture of the human genome,” says Tsunoda.
These findings could also have important ramifications for the conduct of studies into the genetic roots of human disease. Many such investigations are based on so-called ‘genome-wide association studies’ (GWAS), which use known SNVs as starting points for mapping sites in the genome that contribute to the pathology of complex conditions such as diabetes, rheumatoid arthritis or various forms of cancer. However, by over-emphasizing known SNVs, which are by definition more common in the general population, such studies may ignore many rare variants that offer better insight into disease pathology or are more prevalent among select populations, such as individuals of Japanese ancestry.
Tsunoda hopes this work will help steer future population-scale genetic studies as well as the group’s ongoing tumor analysis efforts for the ICGC. “Our findings promote the potential of high-accuracy personal genome sequencing,” says Tsunoda. “We have found that the variations that are functionally relevant to diseases may include lower frequency alleles that are not so common in the population as the SNVs that people are currently using for GWAS, and we may have to sequence individuals' genomes to look at such variations.”
More information: Fujimoto, A., et al. Whole-genome sequencing and comprehensive variant analysis of a Japanese individual using massively parallel sequencing. Nature Genetics 42, 931–936 (2010).