University of Victoria biomedical engineer 'outsmarts' HIV
It is estimated that 38 million people worldwide are currently infected with HIV and that 4.1 million more are added each year. For scientists to design treatment therapies that are effective over the long-term it is essential to learn more about how the virus mutates and develops resistance to medications.
New, groundbreaking research by University of Victoria biomedical engineer Stephanie Willerth has significantly advanced the understanding of HIV and how to treat it.
"The virus mutates at a very high rate which is very problematic for HIV patients because the virus eventually develops resistance to medications,'' explains Willerth, a faculty member with UVic's Department of Mechanical Engineering and the Division of Medical Sciences.
Willerth and her team studied approximately 15,000 different versions of the virus—something that has never been done before. This information has allowed them to locate the specific genes of the virus that were resistant to the drugs—knowledge that could ultimately help researchers develop more effective treatments for HIV.
Willerth says that the methods she used can be applied to other difficult-to-treat viruses such as swine flu, Ebola, influenza or even staph infections.
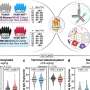
"To study all of these different versions we have to replicate them millions of times, especially when it comes to complex viruses like HIV," explains Willerth. "Because this research method requires a large amount of genetic material and there are obvious risks of duplicating highly contagious viruses, scientists have avoided doing this. Our research was unique because of the method we used—we isolated the genetic material from HIV, so that it was no longer alive, before we replicated it."
After replicating the virus from a small sample obtained from a long-term HIV patient who had developed drug resistance to their treatment, Willerth and her team studied its genetic make-up using "next generation" DNA sequencing—a new method that allows researchers to study millions of molecules at a time.
More information: Willerth SM, Pedro HAM, Pachter L, Humeau LM, Arkin AP, et al. (2010) Development of a Low Bias Method for Characterizing Viral Populations Using Next Generation Sequencing Technology. PLoS ONE 5(10): e13564. doi:10.1371/journal.pone.0013564