New details on how the immune system recognizes influenza
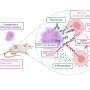
Drawing upon a massive database established with funds from the National Institute of Allergy and Infectious Diseases (NIAID), one of the National Institutes of Health (NIH), scientists have completed the most comprehensive analysis to date of published influenza A virus epitopes--the critical sites on the virus that are recognized by the immune system.
The findings, reported by researchers at the La Jolla Institute for Allergy and Immunology (LIAI), are being published online this week by the journal Proceedings of the National Academy of Sciences.
The study should help scientists who are designing new vaccines, diagnostics and immune-based therapies against seasonal and pandemic influenza because it reveals in molecular detail exactly where the immune system focuses on the viruses. Although the complete molecular structures of essentially all major strains of influenza viruses are known, immune responses concentrate on limited regions of certain parts of the virus, and these regions must be identified as immune epitopes by research studies. The LIAI team found that while there were hundreds of shared epitopes among different virus strains, including the avian H5N1 virus, only one has been published that appears ideal for multi-strain vaccines. Information on shared protective epitopes is important for developing influenza vaccines that can provide broad protection against multiple strains of the virus.
"This study is interesting for what it shows we know and do not know," says NIAID Director Anthony S. Fauci, M.D. "It reveals many gaps in our knowledge of influenza viruses and indicates where we need to focus our attention."
The analysis drew upon a much larger effort called the Immune Epitope Database and Analysis Resources Program, which began in 2004 after NIAID awarded LIAI a $25 million contract to create a single repository of immune epitopes from critical disease-causing microbes, including agents that might be used in a bioterrorist attack. Influenza epitopes comprise only a portion of the extensive database, which has become the largest single collection of such information anywhere in the world. It includes data from thousands of separate articles published over several decades, providing extensive dossiers on dozens of pathogens.
"The purpose of the database is to provide a catalog of molecules and structures that scientists around the world can quickly access and use to understand the immune response to a variety of epitopes, or methodically predict responses to as-yet untested targets," says Alessandro Sette, Ph.D., who heads the Vaccine Discovery division at LIAI and is the lead investigator on the project.
For the current study, Dr. Sette and his colleagues examined 600 different epitopes from 58 different strains of influenza A virus. One of their main goals was to determine how conserved, or similar, epitopes are between different strains of bird and human influenza viruses. Knowing this is important because the virus rapidly mutates and can swap gene segments between strains, which could increase the ability of an avian virus to be transmissible to humans.
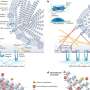
In addition, only a handful of the epitopes are known to be associated with protective immunity. Most of the influenza virus epitopes in the database are those recognized by a type of immune cell known as a T cell; far fewer are recognized by B cells, a type of white blood cell that produces infection-fighting antibodies. Antibodies induced by seasonal and pandemic flu viruses or vaccines are a major component of immunity that protects against these viruses.
Strains of influenza virus can vary enough in their neutralizing B cell epitopes that a vaccine against one strain may not protect against another strain. But if epitopes are conserved between virus strains, the immunity a person has developed towards one strain might provide at least some protection against the other strain.
Using a software tool they developed, the LIAI team found hundreds of conserved influenza virus epitopes in the database, including those between avian H5N1 and strains of human influenza viruses. But what is less clear from the analysis is how cross-reactive an immune response would be to most of these conserved epitopes. Further analyses may assist scientists in identifying vaccine targets that might offer broader protection and in predicting how effective a new vaccine will be.
Other analyses revealed major gaps in scientists' knowledge about influenza viruses. Of the 600 epitopes in the database, for instance, very few were from strains of H5N1 avian influenza. And even though the database contains epitopes from all the influenza virus' proteins, the vast majority of the data relates to just two influenza proteins, the hemagglutinin (HA) and nucleoprotein (NP).
Most of the influenza virus data comes from analyses of immune responses obtained with mice; some comes from rabbits, ferrets and monkeys, and very little comes from humans or birds. In fact, only one antibody epitope came from a human. The LIAI researchers say more studies should be focused on identifying human T and B cell epitopes from human and avian strains of influenza virus--especially those associated with protective immunity.
"The bottom line is that this study shows us where we need to go," says project director Stephen Wilson, Ph.D., chief technology officer at LIAI. "Hundreds of flu epitopes have already been published and are now in the database, but critical gaps become apparent when one looks for human antibody targets."
Plans for the future include adding data on epitopes that are involved in autoimmune diseases and epitopes that trigger allergic and asthmatic reactions. Dr. Sette and his colleagues have also built numerous tools for analyzing and visualizing the data and for predicting immunity against different pathogens--all of which is publicly accessible on their Web site (see http://immuneepitope.org).
Source: NIH/National Institute of Allergy and Infectious Diseases