This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers complete first-ever mapping of human gut virus
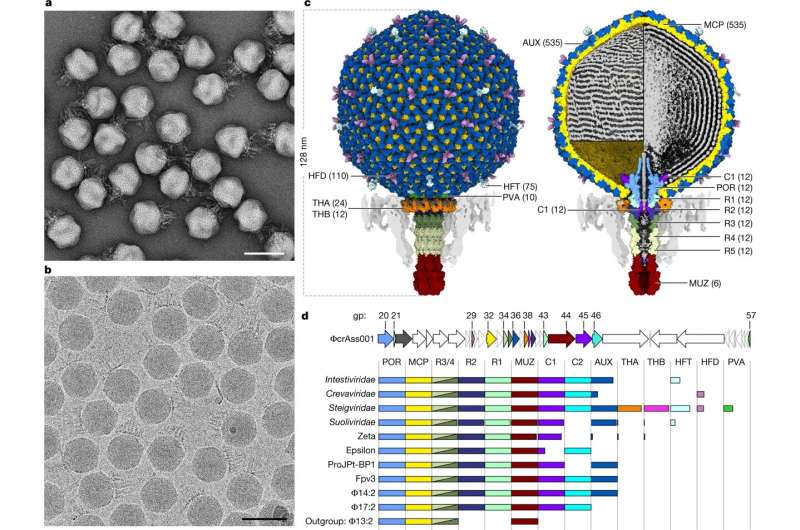
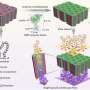
Researchers at APC Microbiome Ireland (APC) have collaborated with scientists in the University of York to complete the first ever structural atlas of a crassvirus (also referred to as crassvirales, bacteriophage, crass-like phage)—the most abundant group of viruses in the human gut, which can play an important role in shaping the gut microbiome, impacting health and disease.
For the first time, scientists have grown a crassvirus in a laboratory and developed a detailed atomic level structure of this gut virus. These gut viruses are believed to play a role in shaping the composition and functionality of the human microbiome. The ability to grow them in the laboratory, together with this newly created structural atlas, will facilitate the research needed to investigate their roles in microbiome functionality and human health. The study is published in the journal Nature.
Key breakthroughs include identifying the presence of multiple cargo proteins carried by the virus, including finding a protein that occupies both the head and tail of the virus. This discovery allows the team to predict a mechanism for how the virus injects its DNA into its bacterial target.
A new protein fold was also identified that acts as a "gatekeeper"—controlling what is transported in and out of the viral particle. Additionally, the team are now able to assign functions to viral genes which were designated as hypothetical until now.
Professor Colin Hill, co-lead study researcher and founding Principal Investigator in APC, said, "These viruses are probably the most abundant biological entities associated with humans, and yet we only found out about them less than a decade ago, so this is all pretty novel and highly exciting."
"Growing the virus is a major research milestone and breakthrough. Essentially, you have to be able to grow it to know it, and once we had developed the ability to grow the virus here at APC, we were able to reach out to colleagues at York to help to study it in all its exquisite detail. The final structure is truly beautiful and is a fitting reward for years of meticulous work at both APC and York," Professor Colin Hill stated.
"This study emphasizes why it is still important to put in the effort needed to study viruses in the laboratory. We can learn a lot from DNA sequencing and computational biology, but you really need to grow a virus in the lab to fully understand it and to inform future experiments," added Andrey N. Shkoporov, co-lead researcher on the study at APC.
Researchers at APC have been studying the gut virome for over a decade and have contributed significantly to knowledge of these highly diverse plethora of viruses. Contained in the gastrointestinal tract, the gut virome varies across human cultures, lifestyles and locations. Professor Shkoporov plans to continue to study the gut virome, including the potential role of bacterial viruses in the spread of antibiotic resistance.
Initially, researchers only knew of the existence of the crassvirus from analyzing the sequences of DNA obtained from gut samples, but a major breakthrough was achieved in 2018 when a team of scientists including Andrey Shkoporov, Professor Colin Hill, Ekaterina Khokhlova and others at APC Microbiome Ireland and the School of Microbiology at UCC grew a member of the crass virus for the very first time. This enabled the research team to collaborate with Professor Fred Antson and Dr. Oliver Bayfield at the University of York to further investigate the virus using cryo-EM (cryo-electron microscopy) technology to comprehensively map the 1,440 proteins that make up the viral particle.
Professor John F. Cryan, UCC Vice President for Research and Innovation and Principal Investigator at APC Microbiome Ireland, said, "Congratulations to Prof. Colin Hill and Prof. Andrey Shkoporov on this latest and important research from APC which provides a detailed structure-function mapping of crassviruses and, combined with comparative sequential analysis, allows us to assign functions to the majority of the previously uncharacterised proteins involved in virion assembly and infection."
More information: Oliver W. Bayfield et al, Structural atlas of a human gut crassvirus, Nature (2023). DOI: 10.1038/s41586-023-06019-2
Journal information: Nature
Provided by University College Cork