This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Biopharming technique yields cost-effective and environmentally friendly antimicrobial peptides

Plants engineered to produce therapeutic peptides could provide a cost-effective and sustainable platform for manufacturing drugs.
As a proof of concept, researchers have coaxed a close relative of tobacco, Nicotiana benthamiana, to churn out peptides with antibiotic activity against some of the nastiest pathogens known to medicine, as others had done in the past.
But, unlike previous efforts to turn plants into drug-production bioreactors, the scientists also modified their shrubs to express a rat enzyme, called PAM, that enhances the stability and prolongs the activity of antimicrobial peptides.
The resulting plants yielded potent drugs that should cost far less to manufacture than those made via other systems—with the added benefit of offering a more environmentally friendly route to drug assembly.
"These plants can be grown on a massive scale, providing a reliable and cost-effective source of medicines for people around the world," says bioengineering professor Magdy Mahfouz, who led the study.
"We now intend to use this technology to produce a wide range of biologics and therapeutics," adds Shahid Chaudhary, a Ph.D. student in Mahfouz's lab group and the first author of the new report.
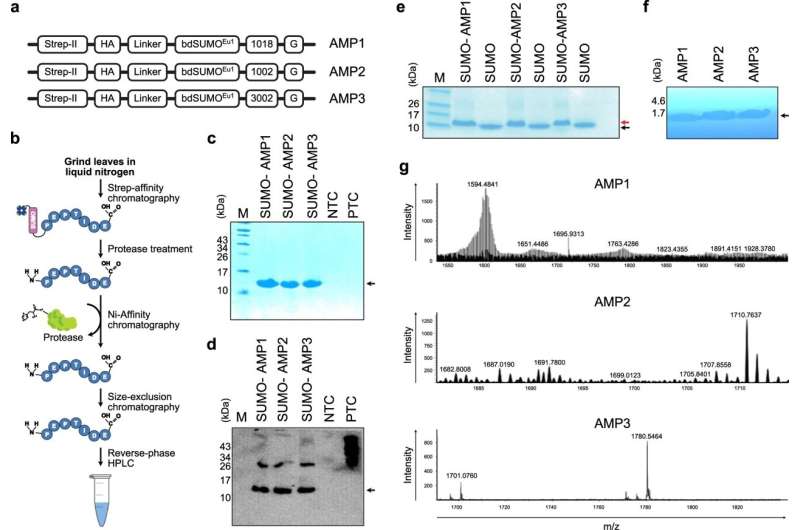
The KAUST research team, which included bioengineers Charlotte Hauser and Samir Hamdan, along with microbiologist Pei-Ying Hong and collaborators from Canada, showed that antimicrobial peptides made in this way could kill several dangerous pathogens, including multiple drug-resistant superbugs responsible for some of the deadliest hospital-acquired infections. The antibiotics also proved harmless to mammalian cells, suggesting that they should be safe for human consumption.
Thinking ahead to eventual deployment of the "biopharming" technique on a massive scale, the researchers showed that their plants were about 3.5-times more efficient at making antibiotics than comparable plants that lack the PAM enzyme modification.
They also added up all the expenses of drug manufacturing and calculated that they could produce 10 milligrams of clinical-grade antimicrobial peptides for less than $0.74 USD—much less than the ~$1000 USD cost of production in commercial companies that chemically synthesize peptides and well below the cost of manufacturing in mammalian cells.
Moreover, plant-based drug manufacturing generates none of the hazardous waste associated with other production platforms, thus making it a much greener option for the pharmaceutical industry.
Mahfouz and his colleagues next plan to make other types of therapeutics in the same way.
"Large-scale industrial production of therapeutic molecules in plants represents a significant step forward in the democratization of medicine," Mahfouz says. "By harnessing the power of molecular biomanufacturing, we can now produce high-quality clinical-grade therapeutics at a fraction of the cost of traditional manufacturing methods."
The study is published in the journal Nature Communications.
More information: Shahid Chaudhary et al, Efficient in planta production of amidated antimicrobial peptides that are active against drug-resistant ESKAPE pathogens, Nature Communications (2023). DOI: 10.1038/s41467-023-37003-z
Journal information: Nature Communications