This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Benchmarking tool capable of closely mimicking single-cell and spatial genomics data
UCLA researchers have developed an "all-in-one," next-generation statistical simulator capable of assimilating a wide range of information to generate realistic synthetic data and provide a benchmarking tool for medical and biological researchers who use advanced technologies to study diseases and potential therapies. Specifically, the new computer-modeling—or "in silico"—system can help researchers evaluate and validate computational methods.
Single-cell RNA sequencing, called single-cell transcriptomics, is the foundation for analyzing genetic makeup (genome-wide gene expression levels) of cells. The introduction of additional "omics" offered detail on a range of molecular features, and in recent years, spatial transcriptomic technologies made it possible to profile gene expression levels with spatial location information of cell "neighborhoods," showing precise locations and movements of cells within tissue.
"Thousands of computational methods have been developed to analyze single-cell and spatial omics data for a variety of tasks, making method benchmarking a pressing challenge for method developers and uses," said Jingyi Jessica Li, Ph.D., a UCLA researcher and professor in statistics, biostatistics, computational medicine, and human genetics. Li is also affiliated with the Gene Regulation research area at the UCLA Jonsson Comprehensive Cancer Center. Li leads a research group titled the Junction of Statistics and Biology.
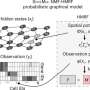
"Although simulators have evolved and become more powerful, there are numerous limitations. Few can generate realistic single-cell RNA sequencing data from continuous cell trajectories by mimicking real data, and most lack the ability to simulate data of multi-omics and spatial transcriptomics. We introduced the scDesign3, which we believe is the most realistic and versatile simulator to date, to fill the gap between researchers' benchmarking needs and the limitations of existing tools," said Li, senior author of a study published May 11 in Nature Biotechnology.
The UCLA researchers say they believe scDesign3 "offers the first probabilistic model that unifies the generation and inference for single-cell and spatial omics data. Equipped with interpretable parameters and a model likelihood, scDesign3 is beyond a versatile simulator and has unique advantages for generating customized in silico data, which can serve as negative and positive controls for computational analysis, and for assessing the goodness-of-fit of inferred cell clusters, trajectories, and spatial locations in an unsupervised way." Goodness-of-fit is a measure of how well a statistical model fits a set of observations.
According to the authors, the system's "transparent modeling and interpretable parameters can help users explore, alter, and simulate data. Overall, scDesign3 is a multi-functional suite for benchmarking computational methods and interpreting single-cell and spatial omics data."
More information: Jingyi Li, scDesign3 generates realistic in silico data for multimodal single-cell and spatial omics, Nature Biotechnology (2023). DOI: 10.1038/s41587-023-01772-1. www.nature.com/articles/s41587-023-01772-1
Journal information: Nature Biotechnology
Provided by University of California, Los Angeles