This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Tracing the evolution of shiitake mushrooms
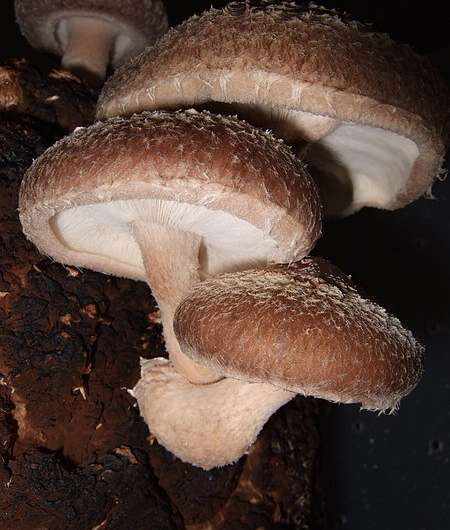

Shiitake mushrooms get their name from the same place they often source their nutrients—the shii tree, a Japanese relative of the oak. These fungi are part of the genus Lentinula, which have evolved to decompose hardwoods on every continent besides Europe and Antarctica.
A new analysis published in the Proceedings of the National Academy of Sciences takes a genomic look at Lentinula samples from around the world. By complete sequencing of 24 mushroom genomes, and assembling genomes from 60 existing sequences, this work expands the family tree for Lentinula. Within those samples, this study also tracks the enzymes these fungi use to break down wood, and compares genetic diversity between cultivated and wild shiitake populations.
Lentinula mushrooms are white rot fungi, belonging to an elite group of decomposers that can break down all of wood's components—cellulose, hemicellulose, and the toughest molecule, lignin. Understanding Lentinula genomes and their evolution could provide strategies for converting plant waste into sugars for biofuel production. Additionally, these fungi play a role in the global carbon cycle. As decayers, Lentinula fungi access carbon previously locked away in wood, converting it into their own carbon-rich structures, as well as atmospheric carbon dioxide.
Researchers sequenced 11 samples from Asia–Australasia and 13 samples from the Americas to trace the evolution of Lentinula species.The Lentinula genus diverged roughly 28 million years ago, when the largest mammal ever, Paraceratherium, still roamed central Asia. Afterward, Lentinula groups evolved further, to four major groups: a lineage in Australasia and L aciculospora, L. raphanica and L boryana in the Americas. Tracing these phylogenetic relationships suggests that the shiitake mushroom, Lentinula edodes, originated in the tropics of the Asia-Australia region. Overall, pangenome analysis showed significantly more genetic diversity in wild Lentinula species compared with farmed cultivars, highlighting a potential conservation threat for wild species.
Sequencing across genomes also provided an opportunity to assess commonalities. The enzymes that Lentinula fungi rely on to decay wood were widely conserved across species. Specifically, many species had very similar plant cell wall degrading enzymes (PCWDEs), including carbohydrate-active enzymes, oxidoreductases, and cytochrome P450s. This includes the species L. raphanica, which has been able to adapt to habitats from Alabama to Brazil, suggesting that the same decay-related enzymes are useful in a wide range of environments.
Collecting and analyzing these samples was a global collaboration that brought together 39 researchers from eight different countries. Genome and transcriptome sequencing were enabled by the Community Science Program (CSP) at the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science User Facility located at Lawrence Berkeley National Laboratory (Berkeley Lab).
More information: Sean Sierra-Patev et al, A global phylogenomic analysis of the shiitake genus Lentinula, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2214076120
Journal information: Proceedings of the National Academy of Sciences
Provided by DOE/Joint Genome Institute