This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Decoding the genetics behind plant height and seed weight scaling in barley
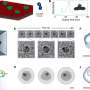
![Correlation of plant height and seed weight [thousand-seed weight (TSW)]. Credit: Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0015 Decoding the genetics behind plant height and seed weight scaling in barley](https://scx1.b-cdn.net/csz/news/800a/2023/decoding-the-genetics.jpg)
Biological functions, resource availability, and evolutionary processes often play a key role in determining the expression of genetic traits and their correlations. In fact, several plant traits are commonly correlated due to different ecological factors. One common example of such correlated traits is plant height and seed size.
This correlation is broadly associated with the allometric scaling (the relative growth rates of different body parts versus the total body size of an organism) following a positive correlation. There are several conceptual frameworks explaining the underlying mechanism of allometric scaling in plants, including a hypothesis that modern crop breeding and domestication can alter the evolution and pattern of expression of plant height and seed weight.
However, the genetic mechanisms influencing size scaling and trait correlation in plants and the actual effects of intense domestication and breeding have remained underexplored.
High-throughput genotyping and large-scale phenotyping hold immense potential to decode the underlying genetic mechanisms of trait correlations and size scaling. Accordingly, a recent study by researchers from Australia have identified the possible genetic factors underlying the correlation between plant height and seed weight scaling in barley crops. The study used genome-wide single-nucleotide polymorphism (SNP) profile for a diverse set of barley samples to measure the differences in plant height and seed weight.
Explaining the rationale for their study, Prof. Chengdao Li, the corresponding author of the study, from Murdoch University says, "Genome-wide association studies (GWASs) have shown that SNPs can help identify genetic variants affecting two or more traits. Moreover, once the physical position of the responsible genetic variant is located on the chromosome, their genetic linkage can be examined."
"We chose barley for our study since it is one of the most important cereal crops that has been subjected to extensive genomic study and has abundant genomic and phenotypic data available."
The study was published in Plant Phenomics.
Prof. Li and his team analyzed the phenotypic data for plant height and seed weight along with the high-density genome-wide SNP dataset for 12,828 barley samples, which included both domesticated and wild barley. They first evaluated the heritability and correlation between both traits, followed by the identification of SNPs associated with either of these traits using GWAS analysis.
Thereafter, they determined the pleiotropic effect (a mechanism for trait correlation where a genetic variant influences two or more phenotypic traits) of each SNP on plant height and seed weight using genomic structural equation modeling. Additionally, they evaluated the effect of genetic linkage on the two traits using linkage disequilibrium (LD) decay.
The researchers found that while plant height and seed weight were heritable and positively correlated in domesticated barley, it was not evident in wild barley. Further, domesticated barley crops are shorter and have smaller seeds than wild barley, which suggests that the plant height and seed weight scaling are less influenced by direct artificial selection.
The team also identified 20 SNPs that caused pleiotropic effects on both plant height and seed weight, and were associated with at least three functional genes related to plant growth and development. Finally, the LD decay analysis showed that a significant number of genetic markers, which influence either or both traits, are closely linked in the chromosome.
These results clearly indicate that two distinct genetic mechanisms form the basis of plant height and seed weight scaling in barley— pleiotropy and genetic linkage.
"Our findings provide a better understanding of the genetic factors underlying size scaling, which, in turn, could expand the scope for exploring other mechanisms underlying the allometric scaling in plants. While our study examined size scaling in a single crop species, it points towards the possible existence of similar genomic mechanisms for allometric scaling across other plant species," remarks Prof. Li.
Moreover, since size scaling in barley is positively correlated, genetically constrained, and affected minimally by domestication, the outcomes of this study could have potential implications for crop breeding where correlated traits are desired in the opposite direction.
More information: Tianhua He et al, Pleiotropy Structures Plant Height and Seed Weight Scaling in Barley despite Long History of Domestication and Breeding Selection, Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0015
Provided by Murdoch University