Mathematical analysis brings new understanding to a recently published kinetic model of ERK phosphorylation
Extracellular Signal Regulated Kinase (ERK) plays an important role in multiple cell signaling processes, by catalyzing the phosphorylation of a variety of substrates. The enzyme itself is activated by phosphorylation at two different sites by a second enzyme called mitogen-activated protein kinase kinase (MEK).
The MEK/ERK pathway is involved in processes such as cell division, differentiation and death, and mutations affecting the MEK/ERK pathway are associated with a range of diseases, including cancer. Understanding the mechanisms and kinetics involved in the MEK/ERK pathway is therefore essential to its exploitation as a therapeutic target.
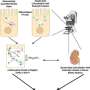
Lewis Marsh from the Byrne lab at the Ludwig Oxford Branch, together with colleagues from the Mathematical Institute and the University of York, undertook a mathematical analysis of a recently published systems biology model by Yeung et al. Their previous analysis of the model relied on assumptions that the Byrne lab mathematically interrogated and confirmed using computational algebra and geometry.
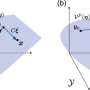
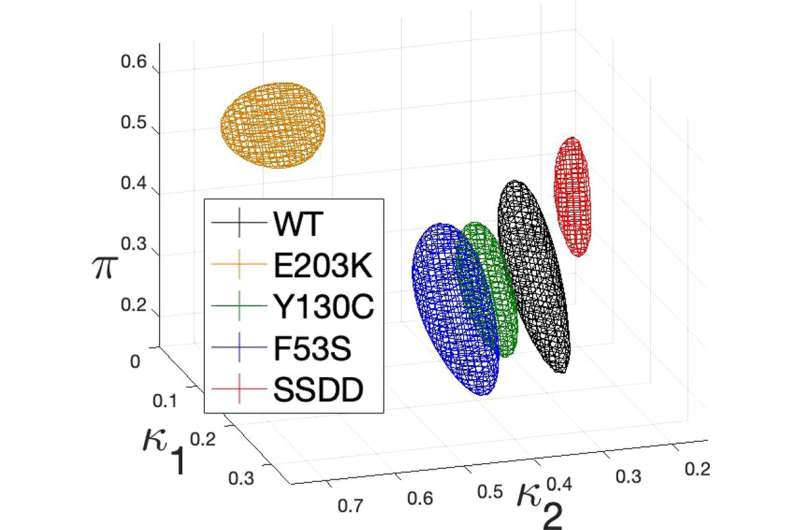
In their study, now published in the Bulletin of Mathematical Biology, Marsh and colleagues analyzed the kinetics of the ERK dual phosphorylation model with both wild-type and pathologically-relevant MEK mutant datasets with topological data analysis. Building on the work of Yeung et. al., this study showcases how algebraic, geometric and topological analysis methods can not only inform and surpass dynamical and statistical analyses of such models, but also guide future experimental design and research directions.
More information: Lewis Marsh et al, Algebra, Geometry and Topology of ERK Kinetics, Bulletin of Mathematical Biology (2022). DOI: 10.1007/s11538-022-01088-2
Provided by Ludwig Cancer Research