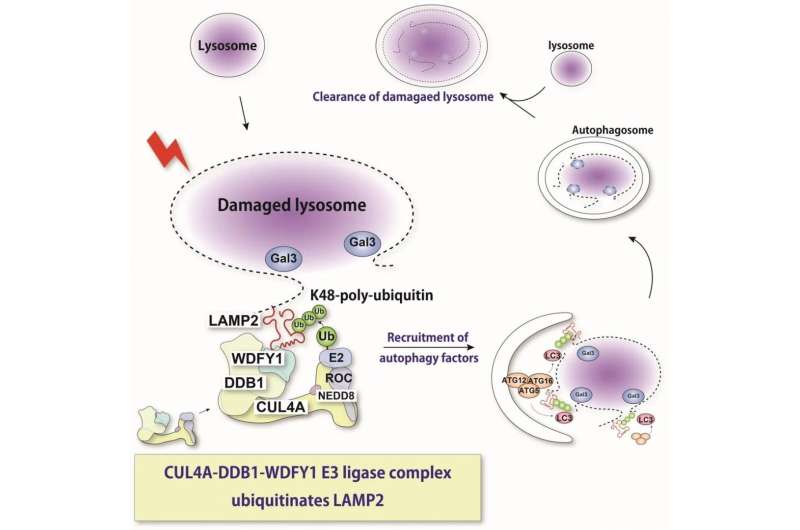
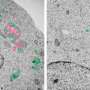
The lysosome is an acidic intracellular organelle crucial for degradation of various cellular components. Upon lysosomal membrane damage, lysosomal hydrolases are released into the cytosol and induce stress responses or cell death. Here, we show that a newly identified protein complex recognizes damaged lysosomes and ubiquitinates a substrate, LAMP2. This recognition induces selective autophagy. Credit: Keisuke Tabata
Autophagy is a self-degradation process that cells use to remove unneeded or damaged components. There are several forms of autophagy, including macroautophagy, which is a bulk degradation system used to target materials in the cell's cytosol to organelles called lysosomes for enzymatic breakdown. However, even lysosomes themselves sometimes need to be degraded. Recently, Osaka University researchers examined the specific molecular details of how damaged lysosomes are selected and marked for clearance.
In a recent article published in Cell Reports, the team described a process called lysophagy, the specific form of selective autophagy responsible for clearing damaged lysosomes. Previous studies have shown that substances like toxins, lipids, and cholesterol or urate crystals can rupture lysosomes. In addition to making the organelle dysfunctional, this damage can also induce oxidative stress and inflammation that may lead to disease development. Therefore, the cell uses lysophagy to address this. However, the mechanisms controlling how cells can recognize the damaged lysosomes and target them for degradation are not fully understood.
"We know from prior investigations that lysosomes can be tagged by a specific enzyme, SCFFBXO27 through a process called polyubiquitination," says one of the lead authors, Hirofumi Teranishi. "Expression of SCFFBXO27 has only been observed in brain and muscle tissues, so we hypothesized that another more ubiquitous enzyme must exist for lysophagy in other cell types."
The team used polystyrene beads coated with a reagent that could induce endosomal damage and then be ubiquitinated. They then isolated the beads through centrifugation and used a method called mass spectrometry to identify the proteins associated with them, ultimately narrowing the list down to 123 proteins.
"With the help of molecular techniques where we could knock down expression of these various proteins, we found that proteins called CUL4A, DDB1, and WDFY1 compose a complex that responds to lysosomal damage," explains Maho Hamasaki, senior author of the study.
Further characterization indicated that this complex acts preferentially during lysophagy and facilitates the addition of the ubiquitin molecules. The WDFY1 protein is needed to specifically recognize the damaged lysosomes.
"We then wondered what part of the lysosome is recognized by this protein complex," says Teranishi. "Numerous lysosomal proteins were examined, until we found LAMP2 to be the one that is ubiquitinated by the CUL4A complex."
The team also found that the presence of LAMP2 and its interaction with WDFY1 are essential for initiating the lysophagy process. Overall, these findings provide critical insights into the molecular mechanisms that are central to lysophagy. This may also help in fighting diseases in which this process is dysregulated. In the future, the researchers plan to determine more precise details about how the CUL4A complex recognizes LAMP2.
More information: Hirofumi Teranishi et al, Identification of CUL4A-DDB1-WDFY1 as an E3 ubiquitin ligase complex involved in initiation of lysophagy, Cell Reports (2022). DOI: 10.1016/j.celrep.2022.111349
Journal information: Cell Reports
Provided by Osaka University