Bacterial intimacy insights could help tackle antimicrobial resistance
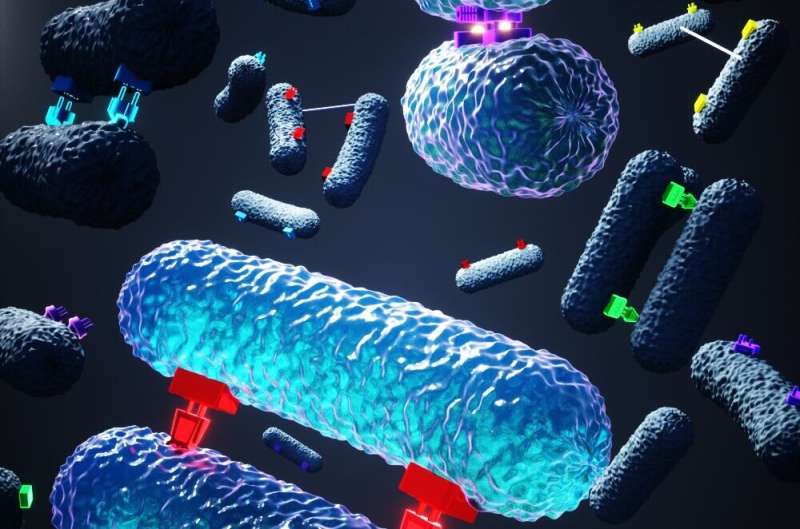
One of the primary ways harmful bacteria acquire resistance to antibiotics is by receiving DNA from other bacteria that are already resistant. This DNA exchange is made via a process called conjugation, akin to bacterial sex, whereby two bacteria form an intimate attachment, and one transfers a packet of DNA to the other.
This is important because antibiotic resistance is causing previously treatable diseases to become deadly. The O'Neill review, commissioned by the UK government, estimates that 10 million deaths could be attributed to infection with resistant bacteria by 2050. Understanding the molecular basis of bacterial conjugation could enable researchers to develop new approaches that slow the spread of antimicrobial resistance.
Since the discovery of bacterial conjugation in the 1940s, much research has been done to show how two bacterial cells initially contact each other in preparation for transfer. However, the mechanism by which donor and recipient bacteria made the intimate attachments that enabled efficient DNA transfer was unknown.
Now, a team led by Imperial College London researchers have uncovered the proteins that mediate these intimate contacts. The results are published today in Nature Microbiology.
The new knowledge could also help scientists predict the spread of emerging resistance amongst bacterial pathogens, as it demonstrates why some DNA packets, called plasmids, are found in specific bacterial species.
Lead researcher Professor Gad Frankel, from the Department of Life Sciences and the MRC Centre for Molecular Bacteriology and Infection at Imperial, said: "The spread of antimicrobial resistance is an acute problem affecting human health globally, and we urgently need new tools to fight it.
"Understanding, and ultimately interrupting, the process by which bacteria share their abilities to evade antimicrobial drugs will go a long way to helping stall the spread of resistance."
Plasmids are packets of DNA that sit inside bacterial cells but replicate separately from the main chromosomal DNA. They carry small numbers of genes that can encode for certain functions, including resistance to antimicrobial drugs.
The team found that during conjugation, a protein from the donor bacteria, termed TraN, acts as a 'plug' to attach itself to a unique outer membrane receptor, or 'socket', in recipient bacteria. Plasmids that are shared by conjugation express one of four variants of the protein TraN, with each variant binding a specific outer membrane receptor in the recipient bacteria, which enables efficient transfer of plasmids from one cell to the other.
The team from Imperial worked with colleagues at the University of Virginia, U.S., to use high-power cryo-electron microscopy to visualize the intimate attachment process, alongside structural biologists also based at Imperial. The group used recent advances in artificial intelligence and bioinformatics to analyse the TraN proteins of several resistance plasmids and the recipient bacteria receptors for several important human bacterial pathogens.
First author Wen Wen Low, from the Department of Life Sciences and the MRC Centre for Molecular Microbiology and Infection at Imperial, said: "These protein-receptor pairings explain conjugation species-specificity. Using plasmid datasets from Enterobacteriaceae—a family of bacteria that include Salmonella and E. coli—we showed how our classification reflects the real-world distribution of resistance plasmids."
Co-author Dr. Konstantinos Beis, from the Department of Life Sciences at Imperial and Research Complex at Harwell, Oxfordshire, added: "These findings present a key advancement in understanding how conjugative mating pairs are formed and will allow us to predict the spread of emerging resistance plasmids into high-risk bacterial pathogens."
The team are continuing to study the interactions of TraN and receptors in detail, including what drives plasmid specialization, and how conjugation dynamics and preferences play out in communities of mixed microbes. They hope that this work will lay the foundations for new approaches to block the spread of antibiotic resistance.
More information: Gad Frankel, Mating pair stabilization mediates bacterial conjugation species specificity, Nature Microbiology (2022). DOI: 10.1038/s41564-022-01146-4. www.nature.com/articles/s41564-022-01146-4
Journal information: Nature Microbiology
Provided by Imperial College London