Simulations predict mysterious biological processes of the cell
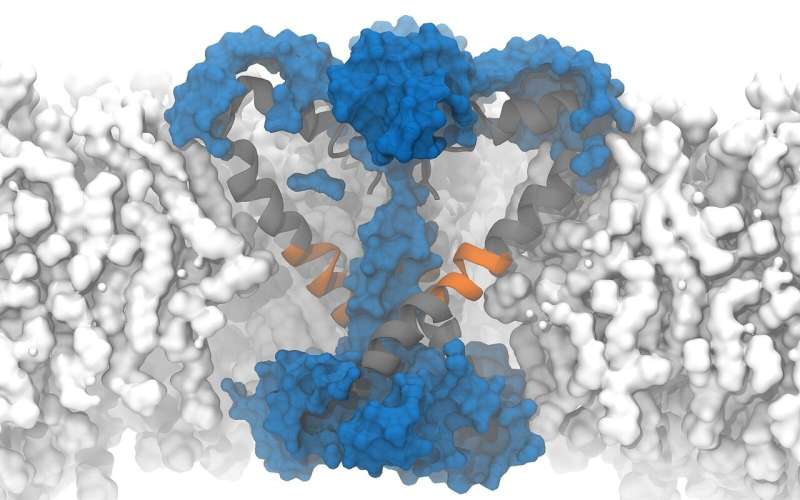
The research group of Lucie Delemotte, Associate Professor in computational biophysics at the KTH Royal Institute of Technology in Stockholm, Sweden, is focusing on understanding the function of ion channels in cell membranes. To do this, the team uses computer simulations on high-performance computers to help model the ion channels formed by protein molecules that span the cell membrane. Voltage-gated ion channels, such as sodium or potassium ion channels, are activated by voltage differences between the extracellular matrix and the cell's cytoplasm. When activated, they open their pores and permit ions to be exchanged between the two sides of the membrane, which triggers electrical signaling.
In muscle or nerve cells—so-called electrically excitable cells—ion transfer causes muscle contraction and enables us to think or dream, explains Delemotte. "If the function of these channels is impaired by genetic mutations, this can lead to epilepsy or muscular paralysis, for example."
In order to be able to treat such diseases efficiently, it is important to know the processes taking place in the ion channels. Better knowledge of how they function helps to personalize the treatment of patients and, above all, to reduce the side effects of drugs.
How exactly these ion channels open their pores to allow ions or active substances to pass through is not yet fully understood. What is clear, however, is that sodium and potassium ions or active substances can only pass through the ion channels if their pores are hydrated. If they are dehydrated, ion exchange is prevented. Until now, it was assumed that pores are hydrated when their diameter is large and dehydrated when the pores are narrow. "In fact, the situation is much more complicated," says the scientist. You can also have a wide pore that is not hydrated if the chemical nature of the pore is hydrophobic.
Key role of water molecules
Scientists experimentally investigate these channels with the help of crystallography or cryo-electron microscopy. This gives us a good idea of how these channels are structured, says Delemotte. However, how the water molecules in the ion channels influence their function cannot be easily observed in experiments. To investigate this, the team used molecular dynamics simulations on Piz Daint. The results were recently published in the Biophysical Journal.
On Piz Daint, a Ph.D. student from the Delemotte team, Koushik Choudhury, simulated the supposedly open structure of an ion channel of a bacterium as it was described based on experiments. Bacterial channels consist of fewer amino acids and are therefore simpler than those of eukaryotes and less complex to simulate. In this bacterium, the channels are equipped with chains of amino acids arranged in a specific structure—so-called α -helix. To the surprise of the researchers, however, the simulation showed that in the model described, the pores are not open and permeable for the ions but rather closed.
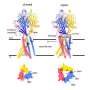
Fortunately, a special circumstance helped the researchers to understand what causes the cell to appear to become permeable: to speed up the simulation, the researchers increased the membrane potential five to 10 times higher than what is found in reality or is possible in experiments. Due to the unusually high voltage field, they accidentally stumbled upon a conformational change of the protein chain that forms the pore of the ion channel. In the simulation, the researchers were able to observe how a kink spontaneously formed in a certain α-helix, which led to the interruption of the hydrogen bonds and the formation of another structure known as a π -helix. The rotation of the end of the protein towards the kink then ultimately resulted in the polar amino acid asparagine at the end of the helix orienting itself towards the pore, and a spontaneous hydration of the pore occurred in a manner compatible with ion conduction.
Following this observation, Delemotte's team then designed a π model. The π helix structure is less stable and can only exist if stabilizing energy is supplied from elsewhere, says Delemotte. And promptly, the researchers' π model showed that the rotation of the π helix with the asparagine towards the pore enables stable ion exchange. They concluded that this process causes hydration and ultimately the opening of the pore.
The protein asparagine has been conserved over the course of evolutionary history and is found in the sodium channels of bacteria, mammals and certain algae. Little is known about its importance, but the fact that it has been conserved suggests that it is an important amino acid, says Delemotte. "Since asparagine is hydrophilic, we think its role in ion channels is to hydrate and, consequently, open the pores, even though we haven't observed this directly in our simulations." Because it is conserved in the sodium channels of all organisms, the researchers hypothesized that such an opening model could also apply to eukaryotic channels—a prediction that would soon prove well-founded.
Just recently, experimental researchers came to a similar conclusion and reported a transformation from a α helix to a π helix in a human channel. Until now, simulations have confirmed experiments; however, "due to the technologies available to us in computational science, we have now reached a point where our simulations are of predictive quality. That is what we have shown with this study," says Delemotte.
More information: Koushik Choudhury et al, An open state of a voltage-gated sodium channel involving a π-helix and conserved pore-facing asparagine, Biophysical Journal (2021). DOI: 10.1016/j.bpj.2021.12.010
Journal information: Biophysical Journal
Provided by Swiss National Supercomputing Centre