Researchers develop new evolutionary approach for identifying proteins that functionally interact
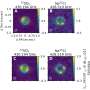
![Extensive long range and interchromosomal gene coevolution. (A) S. cerevisiae and (B) C. albicans differ in chromosome number and size. (C and D) Numbers of genes with interchromosomal orthologous gene coevolution (blue), intrachromosomal (green), or both (orange). (E and F) Intrachromosomal signatures of orthologous gene coevolution corrected by number of genes on chromosome (x axis) and number of interchromosomal signatures of orthologous gene coevolution corrected by number of genes on other chromosomes (y axis). Colors represent different chromosomes, and the regression line of all chromosomes is in black. (G and H) Distances among intrachromosomal signatures of orthologous gene coevolution. (I and J) INO80, an example of how orthologous genes can coevolve with others across the genome. Outermost track: chromosomes of either yeast with chromosome 1 at the 12 o’clock position; second track: genes on plus/minus strand; third track: genes colored according to orthologous gene community. Scatter plot shows the number of coevolving orthologous genes per orthologous gene; size reflects higher values. Links depict orthologous genes coevolving with INO80 and are colored according to chromosomal location of the other orthologous gene. Colors in (E) to (H) and ideogram and link colors in (J) correspond to chromosomes [see (A) and (B)]. Credit: Science Advances (2022). DOI: 10.1126/sciadv.abn0105 Researchers develop new evolutionary approach for identifying proteins that functionally interact](https://scx1.b-cdn.net/csz/news/800a/2022/researchers-develop-ne-7.jpg)
Jacob Steenwyk, a graduate student in biological sciences, and collaborators at Vanderbilt, Tel Aviv University and University of Wisconsin-Madison measured the coevolution of pairs of genes shared across budding yeasts to identify the genes that participate in the same cellular or molecular functions.
The results of this work could fundamentally change how we identify genes with similar functions.
"In this project, we examined nearly 3 million pairs of genes and identified instances where pairs of genes had strong evidence of coevolution," Steenwyk said. "This allowed us to draw a network diagram that reflected cellular and genomic structure and function."
This project shows that genes' evolutionary histories yield insights similar to those of genetic studies on model organisms. This is a hugely important insight because evolutionary analyses are often far less challenging and require fewer resources than genetic studies.
The work was made possible by PhyKIT, a new software co-developed by Steenwyk, and genomic data from hundreds of genomes. PhyKIT is a novel way to use the evolutionary histories of genes to inform our understanding of how proteins interact inside the cell. The target group of organisms used were budding yeasts, a group of microscopic fungi that have a plethora of evolutionary and genetic data available.
"This manuscript is the first major project that heavily leaned on the software I developed," Steenwyk said. "In particular, I used PhyKIT, a broadly applicable toolkit for analyzing and processing multiple sequence alignments and phylogenies."
While Steenwyk remains humble—noting that this area is something of "novel terrain"—his software products have been downloaded more than a hundred thousand times. "Jacob saw a need among evolutionary biologists for user-friendly, robust software. He not only rose to this challenge by developing numerous pieces of software but integrated software development into his approach for asking how the evolution of genes can inform us about their function," Rokas said.
Another interesting result is that gene function, rather than gene location within a chromosome, drives the coevolution of genes. According to the authors, genomes may best be viewed as extensively linked groups of genes. "Our project sets the stage for examining gene coevolution networks in emerging model organisms, lineages with less functional data or lineages that are not yet genetically tractable," Steenwyk said.
The article "An orthologous gene coevolution network provides insight into eukaryotic cellular and genomic structure and function," was published in the journal Science Advances, on May 4.
More information: Jacob L. Steenwyk et al, An orthologous gene coevolution network provides insight into eukaryotic cellular and genomic structure and function, Science Advances (2022). DOI: 10.1126/sciadv.abn0105
Journal information: Science Advances
Provided by Vanderbilt University