Researchers reveal cellular diversity of esophageal tissue
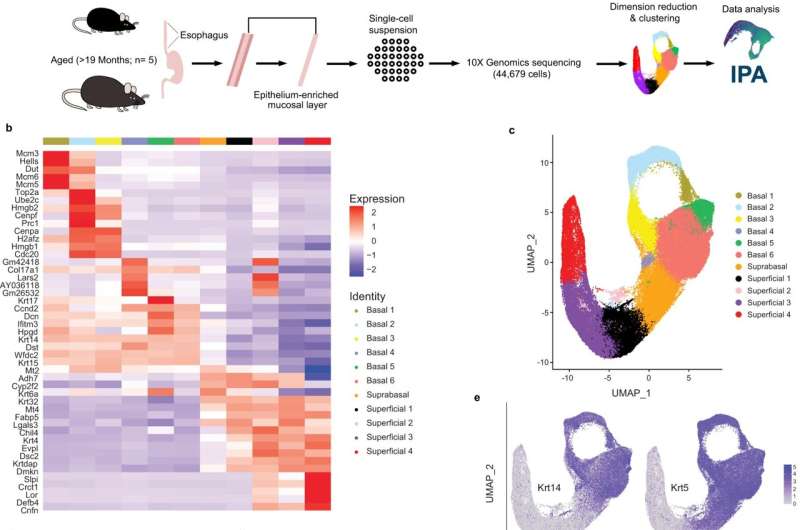
In a study published today in the journal Nature Communications, researchers at the Lewis Katz School of Medicine at Temple University defined 11 subsets of cells found in the esophagus of mice, information that could potentially help clinicians diagnose or treat certain types of cancer.
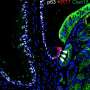
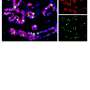
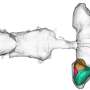
The tissue in the esophagus—known as stratified squamous epithelium—is comprised of a basal layer that ultimately gives rise to several layers of differentiated cells. In this study, researchers sought to find out whether basal cells were all the same or if any differences existed in their gene expressions.
"This work lays the foundation to address questions about how these cells relate to the development of cancer," said Kelly Whelan, PhD, senior author on the study and Assistant Professor of Cancer and Cellular Biology and Assistant Professor in the Fels Cancer Institute for Personalized Medicine at the Lewis Katz School of Medicine.
"The next question we have is if we give a mouse a carcinogen, does one particular subset expand or is another particular subset depleted? If so, we would want to know whether cell types that change in the context of cancer are functionally contributing to the cancer process and whether we can target them to help treat or prevent cancer," she said.
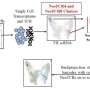
To begin to address these questions, Whelan and her fellow researchers used single cell gene-expression profiling, a tool that identifies all the genes in a cell or tissue that make messenger RNA. These profiles can then be used to identify individual cell types within a complex tissue.
"Essentially, we took more than 40,000 cells from mouse esophageal epithelium and used their individual gene-expression profiles to group them based on their commonality," she said.
When they started the study, Whelan said the researchers expected to find about three subsets of cells classified as basal, but they actually found six subsets, as well as four that were classified as differentiated. Additionally, one cell type was found to have characteristics of both basal and differentiated cells and was defined as an intermediate cell type.
"We are now working to isolate these individual cell types in order to determine what the cells do functionally in terms of esophageal biology in the normal context first and then in the context of cancer. This study is a fundamental step toward that," said Whelan.
More information: Mohammad Faujul Kabir et al, Single cell transcriptomic analysis reveals cellular diversity of murine esophageal epithelium, Nature Communications (2022). DOI: 10.1038/s41467-022-29747-x
Journal information: Nature Communications
Provided by Temple University