Screening study identifies inhibitor of key COVID virus enzyme
When the COVID-19 pandemic hit, scientists across the U.S. Department of Energy's (DOE) national laboratory complex turned to the nation's most powerful supercomputers and other tools to discover molecules that might treat the disease. A study published in the Journal of Chemical Information and Modeling reports the discovery of a molecule with significant potential to disable the virus.
The molecule was identified using high-throughput virtual screening—a search through a library of 6.5 million in-stock compounds that could quickly be scaled up for drug production. The team used computer-based molecular docking studies to identify molecules that could bind to certain targets on the virus's main protease (Mpro)—an enzyme the virus uses to make copies of itself. They also conducted high-throughput laboratory screening experiments, structural studies, and molecular dynamics simulations to learn how these potential inhibitors and the enzyme interact. The goal was to find molecules that could jam up the enzyme's function, which would stop the virus from replicating.
The computational team, which included scientists from the Computational Science Initiative (CSI) at DOE's Brookhaven National Laboratory, identified 72 candidate molecules with potential to inhibit Mpro. Other teams ran laboratory experiments testing those molecules' ability to inhibit the virus. Structural studies using, for example, X-ray crystallography revealed how the candidate molecules fit together with the virus enzyme. Additional computer-based simulations provided details about how those interactions alter the enzyme.
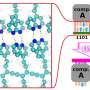
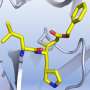
The paper describes how the most promising candidate, known as MCULE-5948770040, binds with Mpro and changes its shape in a way that inhibits the enzyme's function. Future experiments will explore whether the molecule can be developed into a new drug for treating COVID-19.
"Scientists from Brookhaven's CSI played an important role in generating and analyzing large volumes of data that led to scientific insight," said Shantenu Jha, one of the corresponding authors on the paper, who holds a joint appointment with Brookhaven and Rutgers University. "CSI folks also established the 'software infrastructure' to support the large-scale computations," he said.
Given the urgency of the pandemic, "this was a very high-intensity project with a great level of 'learning while doing,'" Jha noted.
CSI's Hubertus Van Dam, another study co-author, agreed, saying he was inspired by "tackling, for me, a new set of problems with a new set of methods in a large team."
The team for this paper included scientists from five national laboratories and four collaborating universities, all supported by the DOE Office of Science through the National Virtual Biotechnology Laboratory (NVBL). NVBL is a consortium of DOE national laboratories focused on response to COVID-19, with funding provided by the Coronavirus CARES Act.
Kerstin Kleese Van Dam, director of CSI at Brookhaven, served as leader of Brookhaven's role in the NVBL medical therapeutics project.
"A key weapon in our arsenal in the fight against COVID-19 are medicines to treat those infected," she said. "The NVBL medical therapeutics project brought to bear the combined might of DOE scientists, and key experimental and computational facilities to discover new COVID treatments.
"This paper describes not only some of our successes, but also gives a glimpse behind the scenes at the scientific ingenuity needed to make those exciting discoveries possible."
More information: Austin Clyde et al, High-Throughput Virtual Screening and Validation of a SARS-CoV-2 Main Protease Noncovalent Inhibitor, Journal of Chemical Information and Modeling (2021). DOI: 10.1021/acs.jcim.1c00851
Journal information: Journal of Chemical Information and Modeling
Provided by Brookhaven National Laboratory