Researchers develop a method that gives enzymes the ability to catalyze new-to-nature reactions

Enzymes are biology's catalytic workhorses, binding molecules together, splitting them apart and reconfiguring them in processes vital to everything from digestion to breathing. Their availability, efficiency and specificity have long made them popular for reactions outside biological systems as well, including those involved in food preservation, detergents and disease diagnostics.
"Enzymes are nature's privileged catalysts," said UC Santa Barbara assistant professor of chemistry Yang Yang. "They can catalyze reactions with amazing selectivity." Efforts over the past three decades have also resulted in the development of customized enzymes—enzymes rapidly evolved for directed purposes, to interact with specific molecules, resulting in a high yield of desired products with unparalleled selectivities.
However, Yang added, the reactions that enzymes can allow for are relatively limited—a somewhat small repertoire for their powerful ability to efficiently make products at lower material, energy and environmental costs.
To bridge that gap and merge the best of both worlds—versatility and selectivity—Yang and his research team have developed a method by which certain enzymes can be coaxed into facilitating useful reactions that were never previously observed in the biological world, thereby widening their repertoire and opening possibilities for processes never before conducted by enzymes.
"If we can develop these so-called new-to-nature enzymatic activities, then we have very powerful biocatalysts for pharmaceutical and agrochemical industries," said Yang, who with colleagues at UC Santa Barbara and University of Pittsburgh authored a paper that appears in the journal Science.
3D chemistry
Stereochemistry (also known as 3D chemistry) is essential to controlling the bioactivity of small-molecule medicines. Most biomacromolecules, including DNAs and proteins, are chiral, meaning they are asymmetric in structure.
"It's just like your left hand and right hand: They look the same but they are not superimposable, meaning they are chiral," Yang explained. "To effectively interact with these chiral biomacromolecules, small-molecule drugs have to be designed with specific stereochemistry. Oftentimes, one enantiomer of a chiral drug molecule is very potent, whereas the other enantiomer is ineffective or even poisonous."
The most efficient way to create such valuable chiral molecules is based on asymmetric catalysis, he said, a process where a tailored catalyst selectively produces one enantiomer (non-superimposable chiral molecule) instead of another. Unfortunately, many challenges still exist in the field of asymmetric catalysis. In particular, a class of widely used reactions—namely radical reactions or reactions involving open-shell intermediates—have not yet succumbed to asymmetric catalysis. This problem has long eluded synthetic chemists.
"Organic radicals are very common and extremely active reaction intermediates in synthetic chemistry," Yang said. "However, it's known that controlling the stereochemistry of these reactions is very, very hard."
There are two problems that arise, he explained. One is that the radical, once generated, usually doesn't interact tightly with the catalyst.
"So there's no way to impose stereocontrol over these radical-mediated bond formations," he said.
Secondly, there is oftentimes the activity-selectivity tradeoff.
"If you have a highly active species, then it's going to be relatively difficult for you to control the selectivity of the reactions involving these intermediates. So there's usually a trade-off," Yang said.
The solution? Directed evolution—evolve the enzyme to be able to rein in the radical.
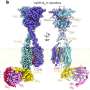
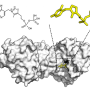
Taking their cue from 2018 Nobel Prize-winning Caltech chemical engineer Frances Arnold, who was Yang's postdoctoral advisor, the team conducted iterative rounds of evolution and screening of cytochromes P450. The superfamily of metalloenzymes is found in all kingdoms of life that contain heme—an iron-containing molecule—that is essential for catalysis.
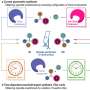
"Directed evolution uses these rounds of mutation and screening to optimize enzyme functions," Yang explained. "In this process, we create a huge library of enzyme variants by DNA manipulation." With a DNA library for target reactions the researchers can express and screen their protein mutants to aid in the identification of promising enzyme catalysts. The improved enzyme then becomes the parent in the subsequent round of engineering. This way, through iterative cycles of mutation and screening, optimal enzyme activity and selectivity are reached.
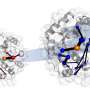
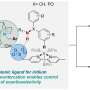
Using this method, the researchers were able to repurpose an enzyme into conducting an "unnatural biocatalytic reaction, namely stereoselective atom transfer radical cyclization," merging the power of synthetic catalysis and control nature affords with enzymatic catalysis.
This new ability opens up many possibilities, including a wider variety of molecules that can be acted upon by newly evolved enzymes.
"The overall goal is to apply the biocatalysts we develop to the pharmaceutical and agrochemical industries," Yang said. "Eventually, with the new tools we'll be able to develop valuable drugs and herbicides which will be useful for our society."
More information: Qi Zhou et al, Stereodivergent atom-transfer radical cyclization by engineered cytochromes P450, Science (2021). DOI: 10.1126/science.abk1603
Journal information: Science
Provided by University of California - Santa Barbara