Artificial intelligence predicts RNA and DNA binding sites to speed up drug discovery
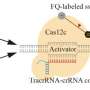
The iMolecule group from Skoltech has developed an artificial intelligence-driven solution that uses data on the structure of RNA or DNA molecules to identify sites on them where interaction with potential drug candidates can occur. Knowledge of these binding sites allows pharmaceutical companies to discover new medications—including antiviral agents—in a much more focused and efficient manner. The new solution is also more accurate than prior approaches, because it accounts for how the shape assumed by a nucleic acid molecule affects which binding sites are exposed. The study came out in Nucleic Acid Research: Genomics and Bioinformatics.
For a long time, pharmacologists have seen RNA as merely a mediator between DNA—that is, our genome—and the functional proteins it encodes, so most drugs target proteins. However, while about 85% of the genome is transcribed into RNAs, only a small fraction of those actually encode proteins. The remaining, noncoding RNAs serve to activate or inactivate certain genes or fulfill other roles by folding into different shapes, called conformations. Since the noncoding functions can take on a pathological dimension, too, RNA and possibly DNA sequences are increasingly recognized as potential drug targets.
"Nucleic acids—DNA and RNA—can participate in signaling, for example, and we could target that or any other process they are involved in. This could be a promising strategy for undruggable protein targets, for example, disordered proteins or proteins that lack convenient binding sites," Skoltech Assistant Professor Petr Popov, the principal investigator of the study, said. "And then there's also pathogenic RNA foreign to the body, for example in viruses, such as SARS-CoV-2 or HIV."
To unlock the potential of all those tentative drug targets, pharmacologists require tools for screening large libraries of chemical compounds to see which of them interact with nucleic acids and what the precise binding spots are.
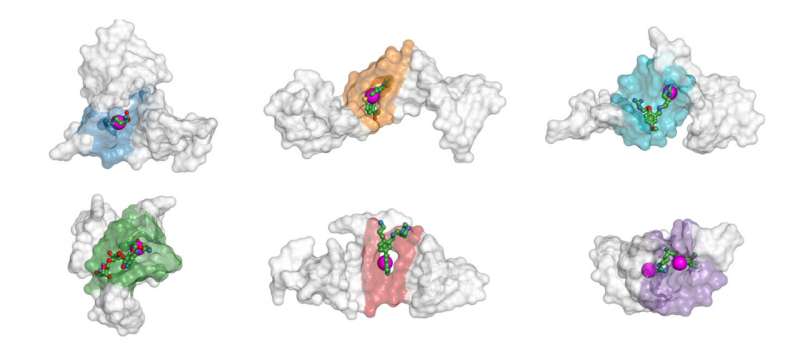
"We created this new solution by adapting our prior work with proteins," Popov explained. "Nucleic acid three-dimensional structures are encoded as high-dimensional tensors. Once this is done, a computer vision algorithm 'looks' at the tensors and highlights the areas in the structure that it thinks could serve as binding sites. After the conformation and the binding site have been detected, a more focused drug discovery campaign can be initiated. So our work is a small step toward rational drug discovery in contrast to the blind screening, which becomes less reliable with growing chemical libraries."
There's an added twist that has to do with the shape of RNA and DNA molecules. They are literally prone to twist and tangle up into distinct shapes. These so-called conformation changes alter the properties of the molecules, including what binding sites are exposed. The conventional approaches only consider the nucleic acid sequences but are blind to conformation and therefore necessarily inaccurate.
"Most earlier methods only worked with RNA, and specifically, with a single chain. Ours works with DNA and with two or more chains. We can even see additional sites that arise when multiple molecules become entangled," Igor Kozlovskii, a Skoltech Ph.D. student and the first author of the paper, said.
"A great example of what makes working with methods that ignore conformation problematic is the dominant type of HIV," he went on. "It has an RNA region targeted by many agents. But even though the nucleic acid sequence is the same, when that molecule changes conformation, this is known to have an effect on which agents work or don't. Our neural network predictions actually reproduce this effect, which means they are reliable."
The new solution has an unexpected application that involves using the method "in reverse." Instead of recognizing binding sites on a potential target, the algorithm could zoom in on a troublesome agent—a small molecule such as a hormone—that is causing a disorder, and distract it.
"So we want to bind those small molecules with something. To do it, we need to reverse-engineer a short nucleic acid fragment, called aptamer, that would serve as a decoy for the hormone or other molecule of interest. Naturally, an aptamer must contain a binding site, and our solution can be applied to design aptamers with improved binding properties," Popov explained.
More information: Igor Kozlovskii et al, Structure-based deep learning for binding site detection in nucleic acid macromolecules, NAR Genomics and Bioinformatics (2021). DOI: 10.1093/nargab/lqab111
Igor Kozlovskii et al, Spatiotemporal identification of druggable binding sites using deep learning, Communications Biology (2020). DOI: 10.1038/s42003-020-01350-0
Journal information: Communications Biology
Provided by Skolkovo Institute of Science and Technology