New tool offers ways to improve CRISPR gene-editing method

The ability to edit the genome by altering the DNA sequence inside a living cell is powerful for research and holds enormous promise for the treatment of diseases. However, existing genome editing technologies frequently result in unwanted mutations or can fail to introduce any changes at all. These problems have kept the field from reaching its full potential.
Now, new research from the laboratory of Princeton University researcher Britt Adamson, conducted with collaborators in the lab of Jonathan Weissman, a member of Whitehead Institute and a professor of biology at the Massachussetts Insittute of Technology and an investigator with the Howard Hughes Medical Institute, and Cecilia Cotta-Ramusino, formerly at Editas Medicine, details a novel method called Repair-seq that reveals in exquisite detail how genome editing tools work.
"We've known for a long time that the mechanisms involved in fixing broken DNA are essential for genome editing because to change the sequence of DNA you first have to break it," said Britt Adamson, senior author on the study and assistant professor in the Princeton Department of Molecular Biology and the Lewis-Sigler Institute of Integrative Genomics. "But those processes are incredibly complex and thus often difficult to untangle."
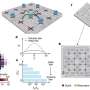
To repair DNA, cells use many different mechanisms, each involving sets of genes working together in distinct pathways. Repair-seq allows researchers to probe the contribution of these pathways to repair of specific DNA lesions by simultaneously profiling how hundreds of individual genes affect mutations produced at damaged sites. The researchers can then generate mechanistic models of DNA repair and learn how those mechanisms impact genome editing. Adamson and colleagues applied their method to one of the most commonly used genome editing approaches, CRISPR-Cas9, which employs the bacterial Cas9 nuclease to cut across both strands of the double-stranded DNA molecule, creating lesions called double-strand breaks.
"Editing with double-strand breaks has been the bread and butter of genome editing for a long time, but making intended changes without unwanted mutations has been an enormous challenge," said the study's first author Jeffrey Hussmann, who conducted the work while a postdoctoral researcher in the laboratory of Jonathan Weissman. "We set out to understand the mechanisms behind as many of the induced mutations as possible, reasoning that this could help us optimize the system."
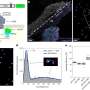
Repair-seq experiments generate an enormous amount of data. Analysis of that data, led by Hussmann, produced a map of how different DNA repair pathways are linked to particular types of Cas9-induced mutations. Building on a rich history of research in the field, Hussmann's analysis illuminated pathways that were already known, and identified new ones, which together highlight the enormous complexity and myriad of systems involved in double-strand break repair. The deep set of data unearthed in this work is now posted on an online portal that others can use to interrogate DNA repair genes and pathways.
Separately, a team led by David Liu at the Broad Institute of MIT and Harvard developed a genome editing system called "prime editing" that doesn't rely on creating double-strand breaks. Prime editing efficiencies vary widely by cell type and target site, but the researchers suspected that identifying the DNA repair pathways involved might help identify avenues for improvement. With this in mind, Adamson and Hussmann joined forces with Liu and colleagues to investigate prime editing using Repair-seq.
"Working together was a huge benefit," said Adamson. "For us, it was a fantastic experience of collaborative and team-oriented science."
The collaborating researchers found that the ability to obtain intended edits with prime editing was affected by proteins in the DNA mismatch repair pathway. They then showed that inhibiting or evading that pathway dramatically enhanced the efficiency and accuracy of prime editing outcomes—positioning prime editing to become a more broadly applicable genome editing technology.
"Working with Britt, Jonathan, and their labs has been a beautiful integration of basic science, tool application, and technology development—a real testament to the power of multidisciplinary collaboration," Liu said.
Importantly, this work also demonstrates how Repair-seq can be used to improve other genome-editing technologies. In fact, the collaborating researchers have already applied it to a third genome editing system, which was also developed by scientists working under Liu. Results from that study were recently published in the journal Nature Biotechnology.
"Repair-seq is a beautiful marriage of technological savvy and biological insight," saidJohn Doench, director of research and development in the Genetic Perturbation Program at the Broad Institute, who was not involved with the work.
"And for the work on prime editing, what a wonderful example of collaboration! Prime editors have often proven difficult to work with, and this paper starts to understand why, while also kickstarting novel solutions," he added.
Moving forward, the team will continue to improve the platform and apply it to additional genome editing technologies.
"We see Repair-seq as a tool that allows you to take a detailed picture of what genome editors are doing inside cells and then very quickly assess, 'Is this a landscape in which I can find design principles that will help improve the tool?'" Adamson said. "We are really excited to explore future applications."
The studies were supported by grants from the National Institutes of Health, the Howard Hughes Medical Institute, the Searle Scholars Program, the National Science Foundation, the Damon Runyon Cancer Research Foundation, the China Scholarship Council, and the National Cancer Institute.
More information: Britt Adamson, Mapping the Genetic Landscape of DNA Double-strand Break Repair, Cell (2021). DOI: 10.1016/j.cell.2021.10.002. www.cell.com/cell/fulltext/S0092-8674(21)01176-4
Journal information: Nature Biotechnology , Cell
Provided by Princeton University