How to program DNA robots to poke and prod cell membranes

Scientists have worked out how to best get DNA to communicate with membranes in our body, paving the way for the creation of 'mini biological computers' in droplets that have potential uses in biosensing and mRNA vaccines.
UNSW's Dr. Matthew Baker and the University of Sydney's Dr. Shelley Wickham co-led the study, published recently in Nucleic Acids Research.
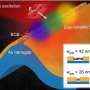
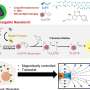
It discovered the best way to design and build DNA "nanostructures" to effectively manipulate synthetic liposomes—tiny bubbles which have traditionally been used to deliver drugs for cancer and other diseases.
But by modifying the shape, porosity and reactivity of liposomes, there are far greater applications, such as building small molecular systems that sense their environment and respond to a signal to release a cargo, such as a drug molecule when it nears its target.
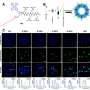
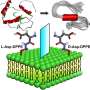
Lead author Dr. Matt Baker from UNSW's School of Biotechnology and Biomolecular Sciences says the study discovered how to build "little blocks" out of DNA and worked out how best to label these blocks with cholesterol to get them to stick to lipids, the main constituents of plant and animal cells.
"One major application of our study is biosensing: you could stick some droplets in a person or patient, as it moves through the body it records local environment, processes this and delivers a result so you can 'read out," the local environment," Dr. Baker says.
Liposome nanotechnology has shot into prominence with the use of liposomes alongside RNA vaccines such as the Pfizer and Moderna COVID-19 vaccines.
"This work shows new ways to corral liposomes into place and then pop them open at just the right time," Dr. Baker says.
"What's better is because they are built from the bottom-up out of individual parts we design, we can easily bolt in and out different components to change the way they work.
Previously scientists struggled to find the right buffer conditions for lipids and liposomes to make sure that their DNA 'computers' actually stuck to liposomes.
They also struggled with the best way to decorate the DNA with cholesterols so that it would not only go to the membrane but stay there as long as was needed.
"Is it better at the edge? The center? Heaps of them? Few of them? Close as possible to structure, or far as possible?" Dr. Baker says.
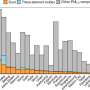
"We looked at all these things and showed that we could make good conditions for DNA structures to bind to liposomes reliably and 'do something.'"
Dr. Baker says membranes are critical in life as they allow compartments to form and therefore different types of tissue and cells to be separated.
"This all relies on membranes being generally quite impermeable," he says.
"Here we have built totally new DNA nanotechnology where we can punch holes in membranes, on demand, to be able to pass important signals across a membrane.
"This is ultimately the basis in life of how cells communicate with each other, and how something useful can be made in one cell and then exported to be used elsewhere."
Alternately, in pathogens, membranes can be disrupted to destroy cells, or viruses can sneak into cells to replicate themselves.
The scientists will next work on how to control DNA-based pores that can be triggered with light to develop synthetic retinas out of entirely novel parts.
More information: Jasleen Kaur Daljit Singh et al, Binding of DNA origami to lipids: maximizing yield and switching via strand displacement, Nucleic Acids Research (2021). DOI: 10.1093/nar/gkab888
Journal information: Nucleic Acids Research
Provided by University of New South Wales