Exploring the genetic ma(i)ze: Functional genomics can help molecular breeding of maize

For centuries, the vast physical diversity of maize, the most popular cereal grain crop globally, has fascinated biologists and plant breeders alike. Early breeders selected the maize variants with good, preferrable traits and propagated them for a long time to develop "true breeding" or inbred lines that show the same favorable features, for instance, the desired kernel color or size, for generations. Because of scientific advances, it is now well known that these trait variations are linked with variations in the genes of maize and that any changes in the "genotype" or gene variation can lead to a new "phenotype" or maize plant variant with a different physical trait.
Finding out which genotype is linked with which phenotype can help us breed plants of desired traits through genetic manipulation, and this process forms the backbone of modern plant breeding. Tissue-culture-based genetic transformation, a commonly used maize breeding technique, however, is limited by two major roadblocks: First, the relative unavailability of genome sequences of inbred lines that show stark phenotypic variations and second, difficulty in inducing callus from immature maize embryos on tissue culture media—the initial step of genetic transformation.
To address these roadblocks, researchers from Sichuan Agricultural University and Berry Genomics Corporation, China, have exploited the power of functional genomics to find solutions to both of these problems. In their study published in The Crop Journal, they explored the genome of maize inbred line A188, known for having many unique traits compared to other lines, to elucidate the linkage between genotype and phenotype. Their choice was based on another advantage, as Dr. Yaou Shen of Sichuan Agricultural University, lead author of the study and one of the corresponding authors, explains: "A188 also shows excellent callus induction from the immature embryo, and therefore, we sought to determine the candidate genes responsible for this highly desirable trait."
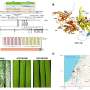
First, they compared the phenotypic variations of A188 with those of three other reference maize inbred lines, B73, Mo17, and W22. Then, they constructed the genomic library of the A188 line. Finally, they annotated the functional aspects associated with the gene products and compared the annotated sequences of A188 with those of the reference lines.
The results turned out promising. The researchers found that A188 had more efficient embryonic callus induction and a relatively higher quality reference genome compared to the other lines. Dr. Langlang Ma, the other corresponding author of the study explains, "We assembled a 2210-Mb genome for A188 with a scaffold N50 size of 11.61 million bases (Mb), a value much larger compared to 9.73 Mb for B73 and 10.2 Mb for Mo17 lines."
Interestingly, almost 30% of the predicted A188 genes were structurally very different from other maize lines, accounting for the high protein divergence and phenotypic variations seen in A188. Comparing the genome sequence with previous data, the researchers identified eight structurally varied genes and two differentially expressed genes as probable key players in maize embryonic callus induction.
Taken these findings together, the researchers surmised that the newly assembled A188 might serve as a new reference genome to clone functional genes of different traits in maize. The study also underscores the importance of A188 to achieve efficient genetic modification of maize through transgenic breeding.
These findings take us one step closer to mapping the genetic maze of maize, and can revolutionize the markets for the world's leading crop.
More information: Fei Ge et al, Genome assembly of the maize inbred line A188 provides a new reference genome for functional genomics, The Crop Journal (2021). DOI: 10.1016/j.cj.2021.08.002
Provided by The Crop Journal