Groundbreaking visualization of atomic movements
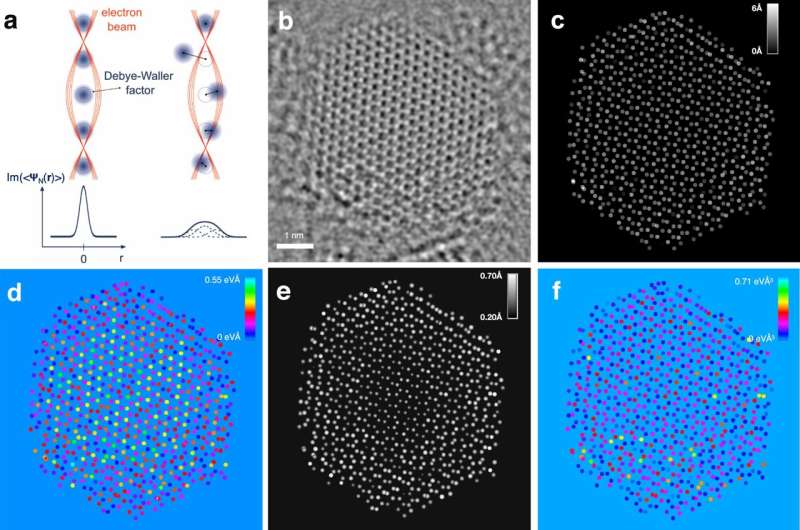
In recent years, a group of leading electron microscopy and catalysis researchers have been working to determine the three-dimensional arrangements of atoms in nanoparticle catalysts in chemical processes. Their work has combined experimental measurements with mathematical modeling.
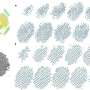
The result is a new method that makes it possible to identify and locate the individual atoms in the nanoparticle, even if they are vibrating and moving.
Until now, atoms in nanoparticles have been expected to be static during observations. But the researchers' analyses of 3D atomic-scale images demonstrated that the original expectation is not sufficient. Instead, the researchers revealed a dynamic behavior of the atoms using a new analytical method.
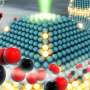
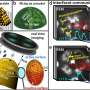
In their work, the researchers have chosen to use a well-known catalytic nanoparticle material, namely molybdenum disulfide. Since the material's atomic structure is well-known, it provided a good basis for interpreting the research group's 3D atomic-resolved images compiled using the unique TEAM 0.5 electron microscope at Lawrence Berkeley National Laboratory, which offers the highest picometre-scale resolution in the world.
The new method is described and published in the renowned scientific journal Nature Communications.
New model ensures the identification of atoms
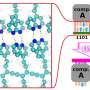
The mathematical model makes it possible to identify the individual atoms in the nanoparticle, even if they are moving. The model measures both the intensity and width of the atoms in the images.
"Until now, determining which atom we are observing has been challenging due to blurring caused by the oscillations of the atoms. However, by factoring in the oscillations, we can more accurately identify, for example, the location of individual sulfur or molybdenum atoms," says Professor Stig Helveg, DTU Physics, who is part of the research group.
The new model also makes it possible to correct alterations of the nanoparticles in the form of oscillations resulting from the illumination of energetic electrons in the electron microscope. It will thus make it possible to focus on the chemical information hidden in the images, atom by atom—which is the essence of the research.
The next step is measuring function
The researchers hope that the new groundbreaking model will find use by other researchers within their field. The model will also provide a basis for Stig Helveg's new basic research center at DTU, VISION.
Here, the focus will be ongoing one step beyond by combining the atomic-resolved images with measurements of the catalytic properties of the nanoparticles. The knowledge produced will contribute to the development of nanoparticles for catalytic processes as part of the transition to sustainable energy.
More information: Fu-Rong Chen et al, Probing atom dynamics of excited Co-Mo-S nanocrystals in 3D, Nature Communications (2021). DOI: 10.1038/s41467-021-24857-4
Journal information: Nature Communications
Provided by Technical University of Denmark