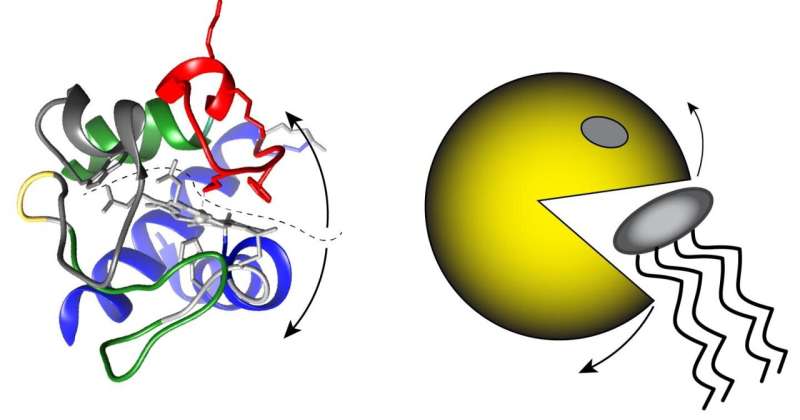
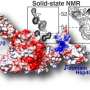
When cytochrome c unfolds, it is able to "attack" cardiolipin and oxidize it, which triggers apoptosis. Credit: Van der Wel lab / University of Groningen
The death of cells is well regulated. If it occurs too much, it can cause degenerative diseases. Too little, and cells can become tumors. Mitochondria, the power plants of cells, play a role in this programmed cell death. Scientists from the University of Groningen (the Netherlands) and the University of Pittsburgh (U.S.) have obtained new insights in how mitochondria receive the signal to self-destruct. Their results were published in the Journal of Molecular Biology.
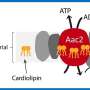
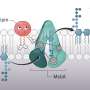
How does a cell kill itself? The details of this process are still unclear. Patrick van der Wel, associate professor of Solid-State NMR Spectroscopy at the University of Groningen, is working together with colleagues at the University of Pittsburgh to see how cell death is initiated on a molecular level. "The membranes of mitochondria play a key role," he explains. Cardiolipin, a special type of membrane lipid, acts as an important signal. "If it gets reshuffled inside the membrane and oxidized, this can trigger cell death."
Unfolding
Another factor is the small protein cytochrome c. This plays an important role in energy production by mitochondria, but it can also bind to cardiolipin. "We believe it may control oxidation of the cardiolipin, which is part of the initiation process for programmed cell death," explains Van der Wel. It was previously thought that the unfolding of cytochrome c on the cell membrane was important, as this allows it to oxidize cardiolipin, a step that triggers cell death. However, in a previous paper, Van der Wel and his U.S. colleagues published evidence that cytochrome c is not unfolded.
"In our new study, we have looked in even more detail at the interaction between cytochrome c and the mitochondrial membrane," says Van der Wel. They used solid-state NMR to detect the position and the status of all 105 amino acids in the protein. The NMR signals of two connected carbon atoms in an amino acid depend on how they interact with other atoms in the molecule. Therefore, the measured spectrum of the carbon atoms can show in which amino acid they are located. Such information can be used to get an idea of the protein's structure, even if it is not well ordered.
Discrepancy
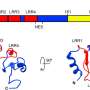
"Furthermore, with this technique, an amino acid is only visible in a fixed part of the protein structure. If it is in an unfolded part, it can move more freely and becomes invisible," explains Van der Wel.Thus, solid-state NMR spectroscopy can show which parts of the protein are folded or unfolded. "What we saw is that cytochrome c is not fully unfolded when attached to the cardiolipins in the mitochondrial membrane."
Proteins fold in a particular order: The first fold induces a second, third, and so on. "These steps are called foldons. What we saw in our experiments is that when binding to the membrane, different foldons in cytochrome c unfold at different stages. And some parts do not unfold at all." This finding explains the discrepancy between the results of Van der Wel and his colleagues and those of previous papers: These studies were too coarse-grained to be able to clearly see which parts of cytochrome c were still folded.
Drugs
These results are interesting, as they provide fundamental insight into how programmed cell death is regulated at the molecular level. "They add to our previous idea that oxidation of cardiolipin by cytochrome c is a very well-controlled, specific process," says Van der Wel. Furthermore, knowing which parts of the protein unfold means that it will be possible to develop drugs that stabilize or destabilize them. Such drugs could be used as regulators that could increase or reduce programmed cell death. Van der Wel continues, "We would like to use our data to construct a realistic computer model for this protein interaction so that it is possible to design these drugs in silico."
More information: Mingyue Li et al, Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids, Journal of Molecular Biology (2021). DOI: 10.1016/j.jmb.2021.167057
Journal information: Journal of Molecular Biology
Provided by University of Groningen