Giant amoeba-infecting viruses rely on DNA-packing proteins
For some giant viruses, a DNA-packing trick may be essential for infectivity.
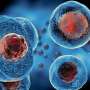
Amoeba-infecting viruses called Marseilleviridae wind their DNA around proteins, like thread loosely wrapped around a spool. This packing arrangement may help Marseilleviridae hijack an amoeba's cellular machinery to replicate itself, reports Howard Hughes Medical Institute Investigator Karolin Luger and colleagues July 22, 2021, in the journal Cell.
The discovery suggests a surprising new role for these DNA-wrapped proteins, called histones, in the life cycle of a virus, says Luger, a structural biologist and biochemist at the University of Colorado Boulder. And along with a recent paper in Nature Structural & Molecular Biology, the new work expands the collection of organisms that have such DNA-packing structures, or nucleosomes.
"As the most exciting science tends to do, this paper opens the door to asking many more questions than it set out to answer," says Tobias Warnecke, a biologist at Imperial College London who was not involved in the study.
Humans, mushrooms, cats, and plants all belong to a group called Eukaryota, organisms that have a defined nucleus surrounded by a membrane. Scientists have known for decades that eukaryotic cells wrap DNA strands around eight histone proteins to form a nucleosome. The prevailing view was that this organization was unique to eukaryotes. But several recent discoveries have challenged this idea.
Scientists have identified histone proteins in microbes called archaea, for example. And Luger's team, among others, has shown that these organisms can bend DNA around histones just like eukaryotes do. Researchers have also discovered histones in Marseilleviridae giant viruses—a type of virus more than twice the size of an influenza virus.
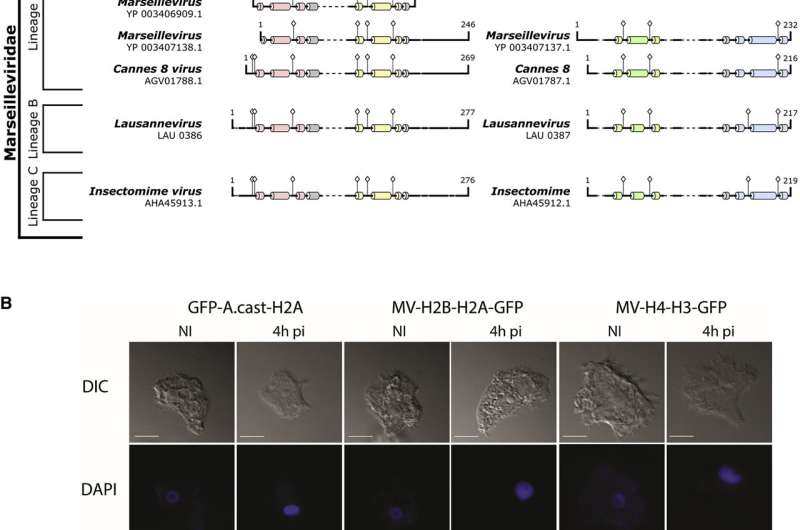
Luger's team wanted to uncover why some giant viruses (sometimes called "giruses") require their own set of histones. The team, which included researchers at the University of Colorado Boulder and Aix-Marseille University in France, conducted in vitro experiments with Melbournevirus, a member of the Marseilleviridae family, to spot clues about the histones' function. The researchers also deleted histone genes from a giant virus and then tested whether it could still infect cells.
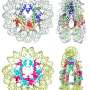
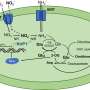
Luger and her colleagues discovered that the histones clumped together in pairs, forming unstable nucleosome-like particles. These wobbly particles have distinct structural and behavioral properties that help the viruses infect their target. Unlike most viruses, which invade a cell's nucleus to replicate, the giant viruses stay in the host's cytoplasm, the watery area outside the nucleus. "It's really weird," Luger says. "There's lots of DNA in the nucleus, so why the heck don't these histones go there to organize the host DNA as well?"
Inside "viral factories" within the cytoplasm of an infected amoeba, the giant viruses package their DNA into bundles. These bundles are less stable and less compact than eukaryotic nucleosomes, the team found. Such loose packing could make the viral genome easy to access, letting host proteins copy the genes needed for the virus to replicate. Without histones, the virus can't organize its DNA and can't successfully infect an amoeba, the team discovered.
Many questions about viral histone proteins remain. Researchers don't know why they're fused in pairs, for instance, or why the virus has nucleosomes in the first place. Other viruses can successfully replicate without the DNA structures, Luger points out.
How giant viruses acquired histone genes and put them to use "is one of those fascinating 'whodunnits' in the evolution of nucleosomes," say Yamini Dalal and Daniël Melters, biochemists at the National Cancer Institute. The new paper from Luger's team enlarges the vast catalog of nucleosome structures spanning the kingdoms of life, they add. "This work unambiguously shows that these viral histones can assemble into nucleosomes very much akin to their eukaryotic counterparts."
More information: Yang Liu et al, Virus-encoded histone doublets are essential and form nucleosome-like structures, Cell (2021). DOI: 10.1016/j.cell.2021.06.032
Journal information: Cell
Provided by Howard Hughes Medical Institute