April 12, 2021 report
Gene sequencing bacteria in natural environment sheds new light on antimicrobial resistance
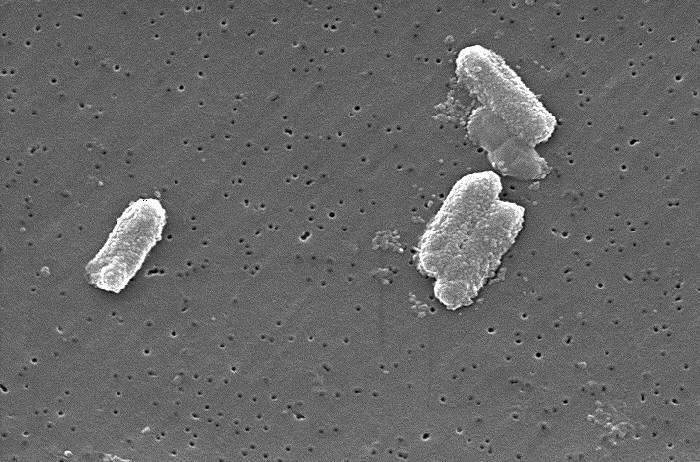
A team of researchers from multiple institutions in the U.K. and the U.S. has learned more about the development of antimicrobial resistance by studying hundreds of samples of bacteria in their natural environments. In their paper published in the journal Science Advances, the group describes how they conducted genome sequencing on hundreds of bacterial samples collected from a wide variety of natural environments and what they learned by doing so.
Over the past decade, medical scientists have grown concerned as more of the kinds of bacteria responsible for infections grow immune to antimicrobial agents meant to kill them. Such antimicrobial resistance (AMR) has become more of a threat in recent times as resistance continues to grow. In this new effort, the researchers have looked at a type of bacteria that are behind intestinal infections. These Enterobacteriaceae include familiar bacteria such as E. coli.
The researchers began their effort by noting that most efforts aimed at better understanding resistance in bacteria have centered around closed environments, such as specimens collected from patients in hospitals. They wondered if more might be learned by taking a look at such bacteria in more natural environments, such as in the soil, in ponds or even in wastewater in treatment plants. To learn more, they collected 2,292 samples from 19 different sites. The samples were then separated allowing the researchers to conduct genome sequencing on 827 kinds of bacteria (553 of which were E. coli).
In looking at their data, the researchers found that some species of bacteria, such as E. coli, that lived in the same environment (such as a pond) shared more genes than had been expected. Notably, they are of a type where genes can move between individual bacterium via horizontal gene transfer. Such sharing was much less likely, they also noted, in more isolated environments. The researchers also found more AMR genes inside of plasmids than in chromosomes. They suggest their findings indicate that Enterobacteriaceae demonstrate both dynamic and plastic AMR gene dissemination and that it is important for researchers involved in AMR efforts to consider natural environments.
The researchers plan to continue their work—they next aim to investigate overlap in environments, including those that involve humans directly.
More information: Liam P. Shaw et al. Niche and local geography shape the pangenome of wastewater- and livestock-associated Enterobacteriaceae, Science Advances (2021). DOI: 10.1126/sciadv.abe3868
Journal information: Science Advances
© 2021 Science X Network