February 26, 2021 report
Scalable software system conducts integrative single-cell chromatin accessibility analysis
A team of researchers from Stanford University, working with associates from the Gladstone Institute of Data Science and Biotechnology and King Abdulaziz University, has developed a software system that can be used for integrative single-cell chromatin accessibility analysis. In their paper published in the journal Nature Genetics, the group describes the software, the platforms on which it can be run, and how it might be used by researchers.
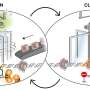
Currently, researchers are able to produce very large datasets containing information about single-cell chromatin, the material that makes up chromosomes, such as DNA and RNA. Unfortunately, tools that allow for analyzing such data are limited. One of the main problems is the amount of additional data that is created as part of such analyses—it grows at such a rate that it overwhelms computer memory. In this new effort, the researchers have overcome that problem with the development of a new software product called ArchR.
The new software is geared toward research involving the way that genes are regulated by the physical arrangement of DNA. ArchR can be used to analyze single-cell data on chromatin accessibility—it shows the ways that DNA can be involved with transcription. More specifically, it can be used to analyze single-cell clustering, double removal, cellular trajectory identification and unified peak set generation.
Making the software even more useful is that it can operate without the need for a high-performance computer. The researchers on the project also claim that it has an easy-to-use interface that for carrying out a variety of analysis scenarios. They also claim that the new software runs some of the same types of analysis schemes as other systems, but faster. And most importantly, because of the way it was designed, memory issues will no longer be a problem. Testing showed it capable of working with much larger datasets than current systems.
The researchers note that the new system provides an easier way to combine transcriptome and epigenome analyses. They suggest that their new software has practical applications for researchers in the field. They next plan to add more functionality to the system.
More information: Jeffrey M. Granja et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis, Nature Genetics (2021). DOI: 10.1038/s41588-021-00790-6
Journal information: Nature Genetics
© 2021 Science X Network