Small differences, big impact: A Hox paradigm for studying protein evolution

In a new study, researchers at the Stowers Institute for Medical Research have identified a handful of variations in an amino acid sequence critical for retaining the ancestral function of a gene over the course of 600 million years of evolution.
The breakthrough discovery detailed in an article published online November 12, 2020, in Genes and Development, offers important insight into the evolution of the gene regulatory networks that drive diversity among organisms, and shows that small differences in key protein sequences can lead to important evolutionary changes.
"It's generally understood that gene duplication and divergence allow genes to take on new functions, while essential genes are conserved and remain unchanged during evolution," says Stowers Investigator Robb Krumlauf, Ph.D. "But how variation in proteins, the building blocks of life, affects this process has been unclear."
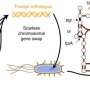
The Krumlauf Lab at the Stowers Institute used a cross-species functional analysis of the labial Hox gene in the fruit fly and related genes in the mouse to explore this question.
The study used modern gene editing technologies, including CRISPR/CAS9, to replace the labial Hox gene in the fruit fly with the three related genes in the mouse—HOXA1, B1, and D1. The researchers found that replacing labial function with HOXA1 in fruit flies restores its original function, but B1 and D1 do not, suggesting that A1 retains an ancestral function, while B1 and D1 have diverged.
"In 600 million years of evolutionary time, only one gene has retained the ancestral activity," says Narendra Pratap Singh, Ph.D., a senior research associate in the Krumlauf Lab and first author of the article. "The other genes evolved and have taken on a new function. This was a great surprise."
The researchers pinpointed a six amino acid sequence critical for the ancestral function of A1, which is important for modulating interactions with other proteins. Also surprising was the fact that the sequence makes up only 2% of total amino acids in the protein, suggesting that tiny differences in certain key regions can have a big impact on protein function.
"Subtle and seemingly innocuous differences in protein sequence can profoundly impact the course of evolution," says Stowers Investigator Kausik Si, Ph.D., an author on the study. "Also, in the evolution of protein function, we tend to focus on what is conserved. This study suggests we should start paying attention to small differences, because some of the most interesting biology is hidden in the tiny differences."
In mice, HOXB1 appears to have evolved to have a new function in vertebrates to allow for greater diversity in facial expression and feeding behavior not found in invertebrates. Mutations in B1 in mice and humans affect facial morphology, neuronal development, and nerve function. In humans, Mobius syndrome, a neurological condition that results in lack of facial expressions, is sometimes associated with B1 mutations.
The study builds on more than three decades of work on Hox genes, a family of "master planner" genes that control the layout of the developing embryo from head to tail. Krumlauf's seminal discovery that Hox genes are essentially the same in mice and fruit flies helped establish the idea that there is a common genetic tool kit and that many organisms have surprisingly similar genes. The lab's comparative studies in mouse, chick, and zebrafish, and more recently sea lamprey, continue to provide critical information on how different species use the same genetic toolkit to form diverse structures. Hox transcription factors are well-suited for investigations into gene duplication and divergence because of their expansion from invertebrates to mammals.
The work paves the way for additional studies on the evolution of protein activity as well as further exploration into the role of conserved toolkit genes following gene duplication and divergence.
"I think we are poised to exploit the emerging strengths of structural biology, functional analyses, and genome engineering," Krumlauf says. "We can really ask, 'Is this role preserved in other invertebrates? Is this gene or protein really doing the same thing or has it evolved completely new functions?' I think there's a new era of analysis now feasible because of the power of gene editing."
Additional contributors to the study include Bony De Kumar, Ph.D., Ariel Paulson, Mark E. Parrish, Ph.D., Ying Zhang, Ph.D., Laurence Florens, Ph.D., and Joan Conaway, Ph.D. This work was funded by the Stowers Institute for Medical Research.
More information: Narendra Pratap Singh et al, A six-amino-acid motif is a major determinant in functional evolution of HOX1 proteins, Genes & Development (2020). DOI: 10.1101/gad.342329.120
Journal information: Genes & Development
Provided by Stowers Institute for Medical Research