Researchers develop simple way to capture high quality 3-D images of live cells and organisms
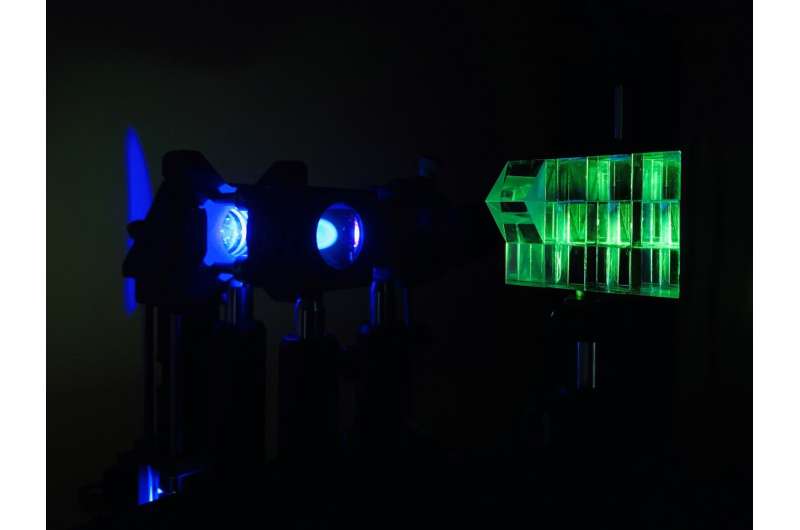

Researchers have developed a simple method for simultaneously acquiring images at different depths with a standard microscope. The new technique can be applied to a variety of microscopy methods, making it useful for a wide range of biological and biomedical imaging applications.
"Optical microscopy has been an indispensable tool for studying 3-D complex biological systems and processes," said Sheng Xiao, a member of the research team from Boston University. "Our new multifocus technique allows live cells and organisms to be observed at high speeds and with high contrast."
In Optica, The Optical Society's (OSA) journal for high impact research, researchers led by Jerome Mertz describe their new straightforward and fast way to acquire information from different depths with standard microscopy. The new approach can be simply added to most existing systems and is easy to replicate, making it accessible to other researchers.
Capturing multifocus images
Standard camera-based microscopy systems acquire sharp images at a single focal plane. Although researchers have tried various strategies to simultaneously acquire images with different focal depths, these approaches typically require multiple cameras or use a specialized diffractive optical element to perform image splitting with a single camera. Both strategies are complex, and a diffractive optical element can be difficult to manufacture.
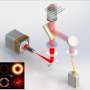
"We used a z-splitter prism that can be assembled entirely from off-the-shelf components and is easily applied to a variety of imaging modalities such as fluorescence, phase-contrast or darkfield imaging," said Xiao.
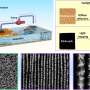
The z-splitter prism divides detected light to simultaneously produce several images in a single camera frame. Each image is focused at a different depth in the sample. Using a high-speed camera with a large sensor area and high pixel count allowed the researchers to distribute multiple high-resolution images on the same sensor without any overlap.
The multifocal images acquired with the new technique make it possible to estimate the out-of-focus background from the sample much more accurately than can be done with a single image. The researchers used this information to develop an improved 3-D deblurring algorithm that eliminates the out-of-focus background light that is often a problem when using widefield microscopy.
"Our extended volume 3-D deblurring algorithm suppresses far-out-of-focus background from sources beyond the imaging volume," said Xiao. "This improves both the image contrast and signal-to-noise ratio, making it particularly beneficial in fluorescence imaging applications involving thick samples."
Demonstrated versatility
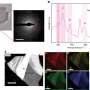
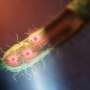
The researchers demonstrated the new technique with commonly used microscopy modalities, including fluorescence, phase-contrast and darkfield imaging. They captured large field-of-view 3-D images encompassing hundreds of neurons or entire freely moving organisms as well as high-speed 3-D images of a rotifer cilia, which beat every hundredth of a second. This showed how the approach provides the flexibility to prioritize a large field-of-view or high speed.
To demonstrate the capabilities of the extended volume 3-D deblurring algorithm, the researchers imaged various thick samples, including the brain of a living mouse. They observed significant contrast and signal-to-noise ratio improvements compared to both raw multifocus images and more traditional 3-D deblurring algorithms. The researchers are now working on expanding the technique so that it will work with even more imaging modalities.
More information: Sheng Xiao et al, High-contrast multifocus microscopy with a single camera and z-splitter prism, Optica (2020). DOI: 10.1364/OPTICA.404678
Journal information: Optica
Provided by The Optical Society