Scientists identify common vulnerabilities in COVID-19 and other lethal coronaviruses
Scientists from the University of Sheffield are working with almost 200 researchers from around the globe to identify vulnerabilities in three lethal coronaviruses—including SARS-CoV-2 responsible for the COVID-19 pandemic.
The international team of experts from 14 leading institutions has studied SARS-CoV-2, SARS-CoV-1 and MERS-CoV to identify commonly hijacked cellular pathways and detect promising targets for broad coronavirus inhibitors with high barriers to resistance. This important research paves the way in identifying a successful treatment for COVID-19.
Using the molecular insights from the study, the researchers also analyzed medical records of approximately 740,000 patients with COVID-19 to examine drugs which are already approved for use and successful in treating other medical conditions and could be deployed rapidly to help the clinical outcomes of these patients.
The findings, published in the journal Science, demonstrate how molecular information can be translated into real-word implications for the treatment of COVID-19, an approach that can ultimately be applied to other diseases in the future.
Dr. Andrew Peden, from the University of Sheffield's Department of Biomedical Science and one of the lead authors, said: "The new insights from this groundbreaking study have revealed potential targets that will help develop a first-of-its-kind therapy across all coronaviruses.
"In Sheffield we were able to bring our broad expertise, as well as the use of our world-class imaging facilities towards a common research goal to desperately find an effective treatment for COVID-19. This study truly demonstrates what can be achieved over a relatively short period of time when scientists openly share ideas, facilities and work for the common good."
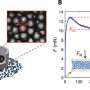
Dr. Peden's team used their expertise in cell biology and advanced microscopy to localize every major viral protein encoded by SARS-CoV-2, SARS-CoV-1 and MERS-CoV inside human cells.
They found that many of the conserved proteins have similar localizations suggesting that they hijack the same cellular processes.
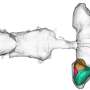
In addition, they also identified that the viral protein Orf9b is localized to mitochondria and alters the levels of Tom70, a key protein which helps cells identify if they have been infected by viruses.
This research, in collaboration with work performed in Freiburg, Paris and San Francisco provides a molecular framework which in the longer term will help in the development of new antiviral therapies which are desperately needed to treat COVID-19.
Dr. Andrew Peden, Dr. Dan Williams and Miss Amber Shun-Shion and are funded by grants from the Biotechnology and Biological Sciences Research Council (BBSRC). All of the imaging studies for this investigation were performed at the Wolfson Light Microscopy Facility at the University of Sheffield.
Professor Colin Bingle from University of Sheffield's Department of Infection, Immunity and Cardiovascular Disease, said: "This systematic cell biology paper used a functional genetic screening approach using real world patient data, to identify potential drugs that can be investigated as treatments for COVID-19. The multinational collaborative nature of this study exemplifies the way that the biomedical community has come together to fight this deadly pandemic."
More information: "Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms," Science (2020). science.sciencemag.org/lookup/ … 1126/science.abe9403
Journal information: Science
Provided by University of Sheffield