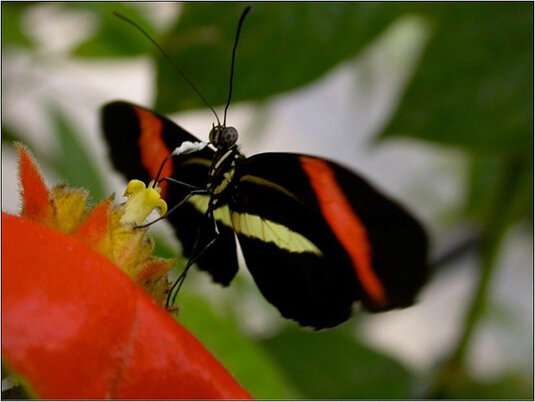
Heliconius melpomene. Credit: R. Merrill
In their efforts to identify the genetic basis for differences in mate choice that keep two co-existing species of butterfly separate, evolutionary biologists at LMU have identified five candidate genes that are associated with divergence in visual mating preferences.
The evolution of a new species often involves a change in mating preference. This happens, for instance, when members of different populations of a given species cease to mate with each other because they no longer find potential partners sufficiently attractive. Two closely related species of tropical butterflies, Heliconius melpomene und Heliconius cydno, provide an interesting example of this phenomenon. The two species are often seen flying together, and crosses between them can result in fertile hybrid offspring.
Nevertheless, individuals of the two species hardly ever mate with each other in the wild. How such behaviorally induced barriers to reproduction emerge is largely unknown. "When changes in behavior are genetically hard-wired, as mate choice seems to be in our butterflies, they must involve alterations in sensation, that is, the stimuli they can detect, or changes in how these stimuli are processed," says LMU evolutionary biologist Dr. Richard Merrill. Together with members of his group, and collaborators at the Smithsonian Tropical Research Institute in Panama and at the University of Cambridge, he has now identified five genes that are linked to the different mating preferences of H. melpomene and H. cydno. As the authors report in the open access journal Nature Communications, these genes are likely to change how visual stimuli are processed during courtship, without altering how the butterflies perceive the world in other contexts.
H. melpomene and H. cydno differ in the striking color patterns of their wings, which serve to warn off potential predators that they are distasteful. H. melpomene has black wings with red bars and thin yellow stripes, while H. cydno's wings feature white bars on a black background. Notably the males of each species show a marked attraction for females with the same 'color scheme' as themselves. In their quest for the genetic factors that underlie differences in these mating preferences, Merrill and his colleagues had previously identified three genomic regions which were associated with the different mating behaviors. One of these regions, on chromosome 18, had an especially strong effect on the degree of persistence with which the male pursues the female of his choice. Strikingly, a gene called optix, which controls the expression of the red bars on the wings of H. melpomene, lies within this same chromosomal segment. However, although this research revealed that one or more genes in this relatively short region of the chromosome must affect mate preference behaviors, the interval in question contains more than 200 genes.
In the new study, Matteo Rossi, a Ph.D. student working in Merrill's group, compared the sequences and activity of these genes in neural tissues—including the central brain, optic structures and the 'ommatidia' (the retinal units that form the facets of the eye) – of H. melpomene and H. cydno. They were able to identify five genes located within this interval that differed between the two species and were associated with their different visual preferences. Most importantly, three of these genes code for proteins that play key roles in neural signal transmission.
"Overall, the nature of our candidate genes suggests that the different preferences for the wing coloration of potential partners are likely to be based on differences in the processing of the visual information. It seems that the two species do not differ with respect to what they see, but they react differently to the different color patterns," says Rossi. "In this way, over the course of evolution, mating preferences can change without affecting perceptions of other aspects of the environment."
More information: Matteo Rossi et al. Visual mate preference evolution during butterfly speciation is linked to neural processing genes, Nature Communications (2020). DOI: 10.1038/s41467-020-18609-z
Journal information: Nature Communications
Provided by Ludwig Maximilian University of Munich