This electron microscopic image of two Epstein Barr Virus virions (viral particles) shows round capsids—protein-encased genetic material—loosely surrounded by the membrane envelope. Credit: DOI: 10.1371/journal.pbio.0030430.g001
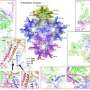
A team of Chinese scientists, led by Prof. Yu Xuekui from the Shanghai Institute of Materia Medica (SIMM) of the Chinese Academy of Sciences (CAS) and Prof. Zeng Musheng from Sun Yat-sen University, reported the first complete atomic model of Epstein-Bar virus (EBV) nucleocapsid. This study was published online in Cell Research.
Like other herpesviruses, EBV, a member of the Gammaherpesvirinae, has a characteristic three-layer configuration: the outer lipid bilayer envelope, the inner nucleocapsid and the middle tegument. The assembly of nucleocapsid is the crucial step in the process of the formation of infectious virion; therefore, revealing the mechanism of EBV nucleocapsid assembly should inform the rational design of anti-viral drugs.
As the first identified oncovirus, EBV is one of the most important human herpesviruses. EBV infects over 90% of the population worldwide and has shown to be closely associated with various malignancies, including Hodgkin's lymphoma, Burkitt's lymphoma, NK/T cell lymphoma and nasopharyngeal carcinoma. In contrast with its medical importance, the structural and functional studies of EBV, largely hindered by the difficulties in sample preparation, are far behind other human herpesviruses, such as human cytomegalovirus (HCMV), herpes simplex virus (HSV) and Kaposi's sarcoma-associated herpesvirus (KSHV).
By developing a novel viral growth method and using cryo-electron microscopy, scientists obtained high quality viral sample of EBV.
Subsequently they determined the high-resolution structure of EBV nucleocapsid and derived its first complete atomic model, including the icosahedral capsid, the capsid-associated tegument complex (CATC) and the dodecameric portal—the viral genome translocation apparatus.
This research described the multi-level interactions between EBV capsid proteins, CATC and capsid, portal and capsid, portal and viral genome, which provided the key structural basis for the development of anti-viral drugs.
In addition, through structural comparisons between EBV nucleocapsid and the counterparts of other herpesviruses, scientists found that the flexibilities of capsid pentons, the copy numbers and the patterns of CATCs binding to the vertices of viral capsid are different among different herpesviruses, and are correlated with the sizes of the packaged genomes.
Based on this observation, they proposed a new pressure-modulating mechanism for how the herpesviruses to achieve a balance between viral genome retention and ejection.
More information: Zhihai Li et al. CryoEM structure of the tegumented capsid of Epstein-Barr virus, Cell Research (2020). DOI: 10.1038/s41422-020-0363-0
Journal information: Cell Research
Provided by Chinese Academy of Sciences