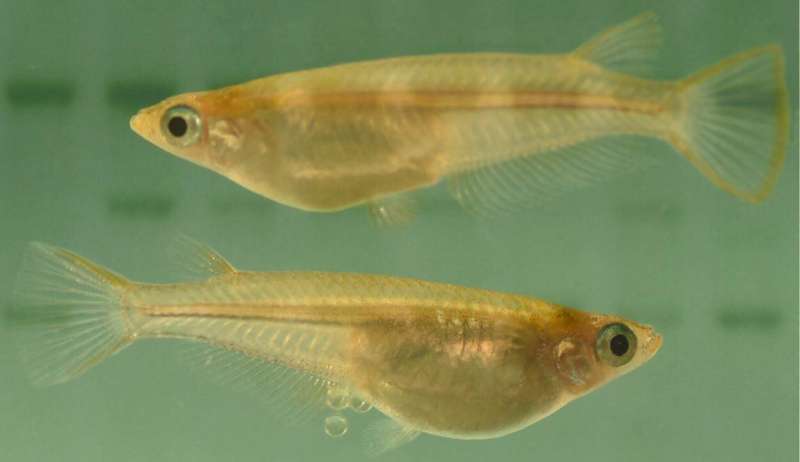
Medaka fish (Oryzias Latipez). Credit: INRAE/Amaury HERPIN
Autophagy is the process by which cells degrade and recycle their own components. It helps maintain homeostasis which is crucial to proper cell functioning. Among the different subtypes of autophagy, chaperone-mediated autophagy (CMA) has been the subject of particular attention in recent years due to the discovery that several human pathologies could be associated to CMA defects. This form of autophagy selectively targets proteins bearing a specific peptide sequence (i.e., the KFERQ motif) present in many protein families. Consequently, CMA helps degrade a wide range of proteins and plays an essential role in the regulation of several cell functions, including gene transcription, DNA repair, cell death and survival, cell metabolism, and the cell cycle. CMA defects have been linked to human diseases, including neurodegenerative diseases, certain types of cancer, metabolic disorders, and immune system diseases.
CMA plays an essential functional role in fish
So far, the absence of any identifiable LAMP2A, a necessary and limiting protein for CMA activity, outside of mammals and birds, led to the paradigm that this cellular function is lacking in other vertebrate lineages. In 2018, French researchers carried out a fine-scale transcriptome analysis in fish using the Phylofish database and discovered that several fish species had transcript sequences displaying high homology with the mammalian version of LAMP2A. This finding suggested that CMA did occur in fish and had appeared much earlier during vertebrate evolution than initially thought (see references below).
In a new study, these same researchers have found clear evidence for this assertion. By using a fluorescent reporter previously used to track CMA in mammalian cells, they revealed the existence of a CMA-like pathway in a fibroblast cell line of the fish medaka (Oryzias latipes). Furthermore, in order to address the physiological role of Lamp2a in fish, they generated medaka knockout for the splice variant lamp2a, and found severe alterations in carbohydrate and fat metabolisms, as previously demonstrated in mice deficient for CMA in liver. Taken together, these results show that CMA is not restricted to birds and mammals: it also occurs in fish, where it plays an essential role in metabolic regulation.
Research implications for a variety of fields
This discovery provides an entirely new understanding of the regulation of metabolism in fish and sheds new light on the evolutionary history of CMA. It offers new perspectives in medical research by enabling the use of complementary genetic models, such as zebrafish or medaka, for studying CMA. Dysfunctions in CMA are known to be involved in several diseases, such as Parkinson's disease, diabetes, immune system disorders, and certain types of cancer.
In aquaculture research, this scientific breakthrough will lead to a better understanding of the mechanisms underlying nutritional modulation of intermediary metabolism, and will provide new metabolic criteria, which can help apply new feeding / breeding strategies in order to optimize fish growth potential while maintaining aquatic product quality and lowering environmental impacts.
More information: Laury Lescat et al. Chaperone-Mediated Autophagy in the light of evolution: insight from fish, Molecular Biology and Evolution (2020). DOI: 10.1093/molbev/msaa127
Journal information: Molecular Biology and Evolution
Provided by INRAE